An in silico Approach for Epitope Based Vaccine Design Against Ebola virus
Article Information
Smrithi Radhakrishnan, M. Elizabeth Sobhia*
Department of Pharmacoinformatics, National Institute of Pharmaceutical Education and Research (NIPER), 160062, Punjab, India
*Corresponding Author: M. Elizabeth Sobhia, Department of Pharmacoinformatics, National Institute of Pharmaceutical Education and Research (NIPER), 160062, Punjab, India
Received: 05 February 2021; Accepted: 16 February 2021; Published: 06 March 2021
Citation:
Smrithi Radhakrishnan, M. Elizabeth Sobhia. An in silico Approach for Epitope Based Vaccine Design Against Ebola virus. Archives of Microbiology & Immunology 5 (2021): 182-206.
View / Download Pdf Share at FacebookAbstract
Ebola viral disease remains a great threat to the society due to the limited availability of licensed drugs or vaccines. As the pathogen belongs to Biosafety Level 4, it is extremely dangerous to design a vaccine using conventional techniques. Therefore, reverse vaccinology approach is used to select the suitable epitopes from the genomic sequence of the virus. Using several algorithms, specific epitopes were chosen and 3D structures were obtained. The HLA alleles vary in different population and hence those alleles that commonly occur in African and world population were selected. Homology models were built for those HLA alleles whose 3D structures were unavailable. Then selected 20 epitopes were docked with selected 8 HLAs and further, selected epitope NQDGLICGL was validated by molecular dynamics study.
Keywords
Zaire ebolavirus (ZEBOV), Glycoprotein (GP); Major Histocompatibility Complex (MHC); Cytotoxic T lymphocytes (CTL); Inhibitory concentration 50 (IC50); Immune Epitope Database (IEDB); Human Leucocyte Antigen (HLA); Auto cross covariance (ACC); Molecular mechanics; The generalized Born model and solvent accessibility method (MMGBSA); Delta G/ Gibbs free energy (DG); Dipalmitoylphosphatidylcholine (DPPC); Three-site transferrable intermolecular potential (TIP3P); Optimized Potential for Liquid Simulations (OPLS); Root mean square deviation (RMSD); Root mean square fluctuation (RMSF)
Zaire ebolavirus (ZEBOV) articles, Glycoprotein (GP) articles; Major Histocompatibility Complex (MHC) articles; Cytotoxic T lymphocytes (CTL) articles; Inhibitory concentration 50 (IC50) articles; Immune Epitope Database (IEDB) articles; Human Leucocyte Antigen (HLA) articles; Auto cross covariance (ACC) articles; Molecular mechanics articles; The generalized Born model and solvent accessibility method (MMGBSA)articles; Delta G/ Gibbs free energy (DG) articles; Dipalmitoylphosphatidylcholine (DPPC) articles; Three-site transferrable intermolecular potential (TIP3P) articles; Optimized Potential for Liquid Simulations (OPLS) articles; Root mean square deviation (RMSD) articles; Root mean square fluctuation (RMSF) articles
Article Details
Introduction
Ebola virus (EBOV) belongs to the filovirus family and is one among the most pathogenic viruses known. It causes a severe hemorrhagic fever in humans and non-human primates having case fatality rates up to 90%. Limited availability of licensed vaccines or therapeutics to treat Ebola viral disease (EBOVD) makes it a very important public health pathogen and the contagiousness together makes it a Category A bio threat pathogen [1]. Infection of humans with filovirus, like Ebola virus and Marburg virus are rare, but the resulting hemorrhagic fevers cause high rate of mortality [2].
Vaccines
Vaccines produce immunity against a given disease-causing pathogen, mostly by inducing the production of neutralizing antibodies against the exposed epitopes. In 2010, a study reported that infectious diseases caused 18.5% of all human deaths and 23% of disability adjusted life years [3,4]. This paved the way for the development of a new method, termed epitope-focused vaccine design. The specific epitopes were selected based on those recognized by naturally occurring antibodies isolated from patients or animal models of a certain disease, aiming to generate protective antibodies in vaccinated individuals to trigger an immune response [5]. This reverse-engineering approach holds promise for the development of vaccines against antigenically diverse viruses [6].
- ChAd3-ZEBOV - The chimpanzee adenovirus – Chimp adenovirus type 3, is used as a vector, to deliver the Ebola genetic material. This vector has a DNA fragment insert encoding the Ebola virus glycoprotein [7].
- rVSV-ZEBOV - The vaccine, uses a replication-competent vesicular stomatitis virus as a vector. It is genetically modified and express surface glycoprotein of the Zaire Ebola species [8].
- Ad26-EBOV and MVA-EBOV – A 2-dose vaccination approach known as the heterologous prime-boost for Ebola has been developed by Johnson & Johnson, in association with Bavarian Nordic. The two vaccine candidates are Ad26-EBOV and MVA-EBOV [9]. The Ad26.ZEBOV is derived from human adenovirus serotype 26 (Ad26) which expresses the Ebola virus Mayinga variant glycoprotein. Meanwhile, MVA-BN is the Modified Vaccinia Virus Ankara - Bavarian Nordic (MVA-BN) Filo-vector [10].
Reverse vaccinology
The use of genomic information with the aid of computer for the preparation of vaccines without the need for culturing microorganism is known as reverse vaccinology, pioneered by Dr. Rino Rappuoli [11]. Since then, this technique has been used for developing several other bacterial vaccines. The genome sequences have the advantage of providing all the protein antigens that the pathogen can express at any time. This approach contains the following steps:
Genome sequences retrieval > Computer analysis > In silico prediction of epitope/ antigen > Development of candidate vaccine in vitro.
The high rate of mortality caused by Neisseria meningitides led to the development of a new in silico technique for vaccine development called reverse vaccinology [12]. The conventional ways for the production of vaccines against meningitis failed because the bacterial and human proteins had a high degree of similarity and also the pathogen had a hypervariable nature. The in silico analysis of the complete genome of Neisseria meningitides helps in specifically selecting the genomic sequences encoding for surface proteins which can act as possible vaccine candidates [13].
The traditional way of vaccine development cannot be employed for the pathogens that are highly infectious and belong to Biosafety Level-4. This led to the discovery of a new approach, to design an efficient vaccine i.e. reverse vaccinology. The major advantage of reverse vaccinology is that, by using bioinformatics techniques a complete genomic sequence of a pathogen can be studied and specific genes coding for potential epitopes can be identified. Preferably for increased antigenicity the genes that code for proteins with extracellular localization, signal peptides and B-cell epitopes are considered [14]. These genes are further screened for desirable attributes that would make it a good vaccine target. The selected genes are then produced synthetically in labs and later screened in animal models [15].
An ideal vaccine is the one which initiates both humoral and cell mediated immunity thus eradicating the chances of re infection [16]. Vaccine design methods based on T cell epitope assist in recognizing immune dominant epitopes of the virus whose synthesis leads to the development of effective vaccine candidates. Computer based prediction methods helps in reducing the number of validation experiments to be carried out for finding the suitable vaccine candidates [17].
Epitope vaccines
An epitope is an antigenic determinant, which helps immune cells in identifying the antigen and has an important role in immunity of an organism. Immune cells recognize epitopes rather than entire antigens [18]. These are seen on the surface of organisms and are detected by antibodies. The selection of epitopes is a crucial step in development of a candidate vaccine. Epitope prediction is the fundamental step in designing of epitope based vaccines [19].
The antigenic peptides are recognized by T cells only when they are presented by MHC I or II, with the help of CD4 and CD8 molecules [20]. T-cell responses have a very important role in controlling viral infections and hence the availability of larger number of T-cell epitope mapping and prediction algorithms is a boon to immunologists [21,22]. For the development of antibody therapeutics [23] peptide based vaccines [23,24] and immunodiagnostic tool, it is very important to accurately identify the B-cell epitopes because a successful peptide-based vaccine must necessarily constitute immune dominant epitopes [25]. Many databases are available based on the kind of epitope, for the large number of epitopes identified in the past 20 years. The databases like MHCPEP, SYFPEITHI [26], FIMM [27], MHCBN [28] and EPIMHC [29] are T cell oriented while Bcipep [30] and Epitome [31] are B cell oriented. AntiJen [32] is a database constituting both T cell and B cell epitopes.
Materials and Methodology
- T cell epitope prediction
VaxiJen 2.0- VaxiJen is a server for alignment-independent prediction of protective antigens. It allows antigen classification solely based on the physicochemical properties of proteins which is a new alignment free approach. It is based on auto cross covariance (ACC) transformation of protein sequences into uniform vectors of principal amino acid properties. The threshold for prediction of viral antigens was set at 0.4 in this server [33].
NetCTL 1.2- NetCTL 1.2 is a server used to predict the CTL epitopes in protein sequences. It integrates the methods of prediction of peptide MHC class I binding, proteasomal C terminal cleavage and TAP transport efficiency. It facilitates the predictions of CTL epitopes restricted to 12 MHC class I supertypes namely A1, A2, A3, A24, A26, B7, B8, B27, B39, B44, B58 and B62. MHC class I binding and proteasomal cleavage is based on artificial neural networks. TAP transport efficiency is predicted using the weight matrix method. The output derived from the neural network predicting MHC/peptide binding is a log transformed value related to the IC50 values in nM units. In this server, scores from all the three individual prediction methods are integrated as a weighted sum with a relative weight on peptide/MHC binding of 1 [34].
CTLPred- The identification of peptides that can stimulate cytotoxic T lymphocytes (CTLs) is one of the major challenges in the design of subunit vaccines. The server employs machine learning techniques Support Vector Machine and Artificial Neural Network to develop a CTL epitope prediction method. An ANN (Artificial Neural Network) based method has been developed for the classification of the large dataset of CTL epitopes (1137) and non-epitopes (1134).
SYFPETHI- The increasing number of motifs necessitated the setting up of a database which facilitated the search for peptides and allowed the prediction of T-cell epitopes. The prediction is solely depended on the published motifs (pool sequencing, natural ligands) and it takes into consideration the amino acids in the anchor and auxiliary anchor positions, as well as other frequent amino acids. The score is calculated as per the following rules: the amino acids of the given peptide are given a specific value depending on whether they are anchor, auxiliary anchor or preferred residue. 10 points are set for ideal anchors, 6-8 points for unusual anchors, 4-6 points for auxiliary anchors and 1-4 points for preferred anchors. Negative values between -1 and -3 are set for amino acids that are regarded as having a negative effect on the binding ability [35].
IEDB analysis resource t-cell epitope prediction tool- This tool enables the prediction of IC50 values for peptides binding to specific MHC molecules. It will take in an amino acid sequence, or set of sequences and determine the ability of each subsequence to bind to a specific MHC class I molecule. The underlying principles behind this tool are Artificial Neural Network, ARB matrix application, Stabilized Matrix Based Method, NetMHCcons and PickPocket method.
T-cell epitope processing score prediction- This tool predicts the epitope candidates on the basis of the processing of peptides in the cell. It combines prediction algorithms of proteasomal processing, TAP transport and MHC binding to produce an overall score for each peptide's intrinsic potential of being a potential T cell epitope.
IEDB analysis resource tool immunogenicity predictor- This tool uses amino acid properties and their position within the peptide in order to predict the immunogenicity of a peptide MHC (pMHC) complex.
AllerTOP v.2 server- The server works on the basis of auto cross covariance (ACC) transformation of protein sequences into uniform equal-length vectors. ACC is a method for protein sequence mining used for quantitative structure-activity relationship (QSAR) studies of peptides with different lengths. The principal properties of the amino acids are indicated by five E descriptors used to describe amino acid hydrophobicity, molecular size, helix-forming propensity, relative abundance of amino acids and β-strand forming propensity. The proteins are classified by k-nearest neighbor algorithm (kNN,k=1) and it is based on training set which contains 2427 known allergens from different species and 2427 non-allergens [36].
Epitope prediction
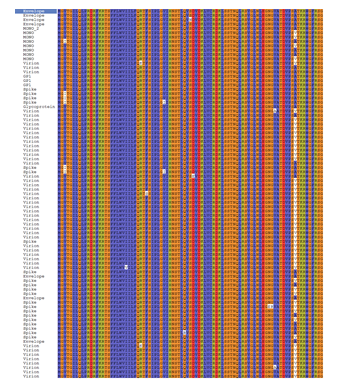
The EBOV glycoprotein sequences available in the UniProtKB database were retrieved. The incomplete sequences were filtered out and only complete sequences were submitted to Maestro for conservancy analysis. These sequences of ZEBOV GP were found to have more than 90% sequence similarity and hence highly conserved. The ZEBOV GP of the 1976 outbreak, Mayinga strain was selected for further studies due to the availability of its crystal structure.
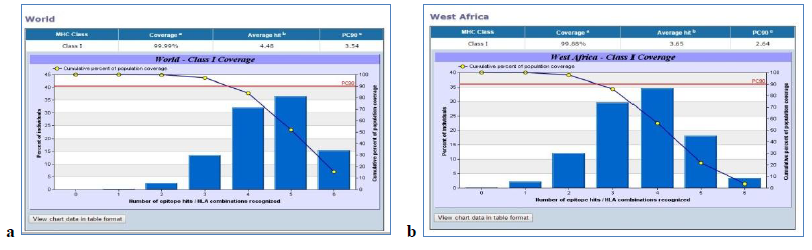
The VaxiJen server predicted the antigenicity score of the glycoprotein as 0.4929 which marked it as a probable antigen. NetCTL 1.2, SYFPETHI and CTLPred servers were used to predict the T-cell epitope and the epitopes in common were selected. The IC50 values were found using IEDB Analysis Resource T-cell Epitope Prediction Tool for the selected epitopes and those epitopes which had less than 200nM value were considered for further analysis. The processing scores were calculated using T-Cell Epitope Processing Score Prediction and 0 was considered the cut off. The epitopes having scores greater than 0 were screened. Using IEDB Analysis Resource Tool conservancy analysis were done and the epitopes more than 90% conserved were selected. IEDB AR Tool was used for immunogenicity prediction and the Population Coverage Tool for analysis of population coverage. The HLA alleles were selected by using the dbMHC data and IMGT/HLA database. Different sets of HLA alleles were given for African population and world population. The population coverage for North, West, East, South and Central African population was found to be 99.84 when the population as a whole was selected and 66.37 when each population were selected individually. In case of world population 99.99 and 65.91 were the values obtained.
Allergenicity of the epitopes were predicted using AllerTop v2.0 and non-allergenic epitopes were further processed. Using Pymol the epitopes were visualized and the PDB resolved portion was isolated. RTFSILNRK, FQRTFSIPL, RTSFFLWVI, STHNTPVYK, RTFSIPLGV, FAFHKEGAF, GYYSTTIRY, VIALFCICK, NQDGLICGL, AEGVVAFLI, GTNETEYLF, FFLWVIILF, TTIGEWAFW, FAEGVVAFL, YYSTTIRYQ, TSFFLWVII, SFFLWVIIL, FLWVIILFQ, YEAGEWAEN and TFAEGVVAF are the epitopes whose structures were elucidated. These were submitted to docking study. The sequence of the epitopes present in the unresolved region of PDB structure were submitted to PEPstr server.
- Allele Preparation
Homology modeling
Knowledge of the three-dimensional structure is necessary for rational drug design, studying effect of various parts of protein on substrate/inhibitor binding etc. The sequences of Human HLA-A*23:01, HLA-A*30:02, HLA-B*15:03 and HLA-Cw*07:02 were retrieved. The alignments were done with HLA-A*24:02(PDB ID: 3I6L), HLA-A*03:01 (PDB ID: 2XPG) and HLA-B*15:01 (PDB ID: 1XR8) respectively, using Modeler 9.14. Multiple template alignment was carried out for HLA- Cw*07:02 using HLA-Cw*04:01 (PDB ID: 1QQD), HLA- Cw*03:04 (PDB ID: 1EFX) and HLA-C*08:01 (PDB ID: 4NT6). Comparative modeling was then performed using MODELLER 9.15 and PyMol. Analysis of models were performed using UCLA‘s SAVES server.
Homology modeling consists of following steps:-
Template selection > Target-Template alignment (BLAST) > 3D structure Modeling (MODELLER 9.15 and Pymol) > Model validation (RMSD, Ramachandran plot, Rampage and Errat plot) > Model refinement
Template selection
The commonly occurring HLA alleles in humans with main focus on the African population were selected and sequences were retrieved from UniProtKB database. The PDB structures of 4 HLA alleles were not found in PDB namely HLA-A*23:01, HLA-A*30:02, HLA-B*15:03 and HLA-Cw*07:02 and they were subjected to homology modeling. Their sequences were subjected to sequence similarity search using BLAST against Protein Data Bank. As a result, many hits were obtained and the ones having better identity/similarity were selected. HLA-A*24:02(PDB ID: 3I6L), HLA-A*03:01 (PDB ID: 2XPG), HLA-B*15:01 (PDB ID: 1XR8) and HLA-C*08:01 (PDB ID: 4NT6). It was found that the HLA had a heterodimer crystal structure. The percentage identity were found to be 99%, 97%, 99% and 94% respectively.
Target-Template alignment
The homology models of the HLAs were built by using the alignment shown below. Stars indicate identical amino acids while colon for similar amino acids and single dot for nearly similar ones.
3D structure modelling
Homology modeling was performed as follows
The sequences of Human HLA-A*23:01, HLA-A*30:02, HLA-B*15:03 and HLA-Cw*07:02 were retrieved. The alignments were done with HLA-A*24:02 (PDB ID: 3I6L), HLA-A*03:01 (PDB ID: 2XPG) and HLA-B*15:01 (PDB ID: 1XR8) respectively, using Modeler 9.14. Multiple template alignment was carried out for HLA- Cw*07:02 using HLA-Cw*04:01 (PDB ID:1QQD), HLA- Cw*03:04 (PDB ID: 1EFX) and HLA-C*08:01 (PDB ID: 4NT6). Comparative modeling was then performed using MODELLER 9.15 and PyMol. Analysis of models were performed using UCLA‘s SAVES server.
Model validation was done using Ramachandran plot and Errat plot.
- Computational details
Molecular docking
BioLuminate is one of the widely used peptide docking programs and it uses a series of hierarchical filters to search for possible locations in the active site region of the receptor. The properties of a receptor/active site region is represented by a grid possessing different set of fields that provide progressively more accurate scoring of the ligand pose. It uses a Glide score (Gscore) and MMGBSA score for predicting binding affinity and rank ordering of ligands in the database screening [37].
Peptide docking consists of the following steps:
Protein preparation – The macromolecules were processed, refined and minimized with the Protein Preparation Wizard at pH 7.0. Water molecules with less than 3 H bonds to non-waters were removed. Other parameters were set default. OPLS_2005 force field was used throughout docking.
Ligand preparation The 20 epitopes were subjected to peptide capping and minimized after adding hydrogen. Epik and Confgen were used for generating conformers of ligands.
Peptide docking was performed for twenty ligands against eight macromolecules with BioLuminate.
The grid box was generated surrounding selected residues. The size of grid box was 20Å in all three dimensions.
Table 1: Protein with surrounding residues in the grid box
|
Protein |
Residues |
|
HLA-B*53:01 (PDB ID:1A1M) |
Asp 114, Tyr 159, Trp 147, Asn 77, Tyr 171, Asn 77, Tyr 171, Tyr7, Arg 62, Tyr 74, Tyr 84, Ile 80, Lys 146, Tyr 99, Arg 97, Asn 63, Tyr 59, Tyr 123, Tyr 59, Ile 66, Phe 67, Tyr 9, Asn 70, Met 5 |
|
HLA-B*51:01 (PDB ID:1E27) |
Tyr 159, Tyr 9, Asn 70, Thr 73, Tyr 7, Asn 77, Trp 147, Tyr 84, Thr 143, Trp 95, Trp 167, Tyr 59, Arg 62, Asn 63, Ile 66, Phe 67, Tyr 99, Tyr 74, Tyr 116, Glu 152, Lys 146, Ala 81, Tyr 123, Tyr 116, Glu 76 |
|
HLA-B*35:01 (PDB ID:3LKN) |
Tyr 84, Leu 81, Tyr 123, Thr 143, Lys 146, Asn 80, Ser 77, Trp 147, Thr 73, Thr 69, Gln 55, Tyr 99, Tyr 159, Tyr 7, Tyr 171, Tyr 59, Ser 116, Asn 70, Leu 156, Ile 66, Asn63, Phe 67, Leu 163, Arg 62, Phe 33 |
|
HLA-A*02:01 (PDB ID:3MRE) |
Tyr 171, Tyr 59, Tyr 7, Glu 63, Tyr 99, Thr 99, Trp 147, Tyr 84, Lys 146, Thr 143, Tyr 123, Trp 167, Met 5, Leu 81, Val 76, Asp 77, Thr 80, Val 152, Gln 155, Leu 156, Arg 97, His 70 |
|
HLA-A*30:02 |
Trp 84, Pro 81, Tyr 123, Asp 143, Asp 146, Arg 80, Glu 77, Tyr 147, Ala 73, Met 69, Ser 116, Ser 95, Glu 70, Ala 163, Ser 33, Arg 99, Ala 159, Trp 171, Arg 59, Gln 179, Leu 184, Arg 194, Gly 50, Thr 104, Tyr 108, Gln 120 |
|
HLA-A*23:01 |
His 138, Arg 30, Ala 141, Gln 179, Ala 182, Thr 187, Gly 191, Trp 84, Pro 81, Phe 123, Asp 143, Asp 146, Gly 80, Glu 77, Tyr 147, Ser 95, Ser 156, Ala 163, Ser 33, Arg 155, Arg 99, Ala 159, Trp 171, Arg 59 |
|
HLA-B*15:03 |
Trp 84, Pro 81, Tyr 123, Asp 143, Asp 146, Gly 80, Glu 77, Tyr 147, Ala 73, Ser 116, Thr 95, Ala 70, Ala 163, Tyr 33, Tyr 51, Arg 181, Val 189, Arg 194, Val 49, Arg 99, Trp 171, Arg 59 |
|
HLA-Cw*07:02 |
Trp 84, Pro 81, Ser 123, Asp 143, Asp 146, Gly 80, Glu 77, Tyr 147, Ala 73, Gly 69, Ser 116, Ala 95, Glu 70, Ser 156, Ala 163, Asp 33, Arg 99, Arg 155, Ala 159, Leu 171, Arg 59, Tyr 195, Thr 187, Leu 102, Gln 179 |
Peptide docking box dimensions were set automatically based on peptide ligand set. Default parameters were used. Number of poses to return for each independent docking run was set to 1 for all macromolecules except HLA-B*51:01, for which it was 10. Scoring method used was MM-GBSA which is more accurate than Glide score which is faster.
The Glide score or docking score is an estimate of the binding affinity, but it is only accurate to a few kcal/mol. Its use is limited to distinguishing actives from inactives. MMGBSA allows receptor flexibility and is based on molecular mechanics applications. It is more accurate than Glide score and is usually set as criteria in peptide docking.
Molecular dynamics
Molecular dynamics was carried out in Desmond package of Maestro 9.7. As HLAs are membrane proteins, DPPC membrane model was set up at residues between 136 and 176 of protein chain for HLA-B*53:01 and between 196 and 166 of protein chain for HLA-Cw*07:02. TIP3P water was the solvent model used with orthorhombic boundary of size 10X10X10 Å. Salt concentration used was 0.15M with positive ion Na+ and negative ion Cl-. Surface tension was 4000 barÅ. Box size calculation method was opted as buffer. Other default parameters were chosen. OPLS_2005 force field was used.
Simulation was carried out for 10ns. NPT ensemble class was used with temperature 300K and pressure 1.01325 bar. The model system was relaxed before simulation.
Results
Homology modeling
Model validation
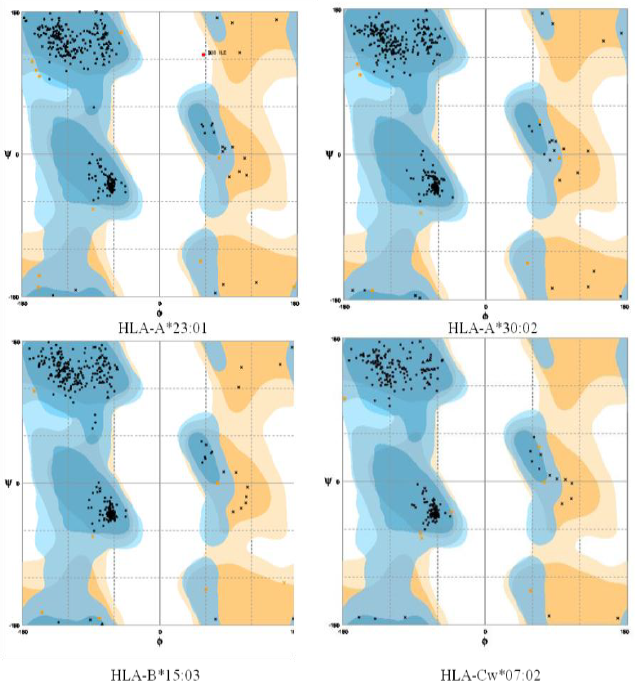
Ramachandran plot- The Ramachandran plot displays the psi and phi backbone conformational angles for each residue in a protein. In a Ramachandran plot, the core or allowed regions are the areas in the plot that show the preferred regions for psi/phi angle pairs for residues in a protein. If the determination of protein structure is reliable, most pairs will be in the favored regions of the plot and only a few will be in disallowed regions. The Ramachandran plot analysis was done by RAMPAGE server.
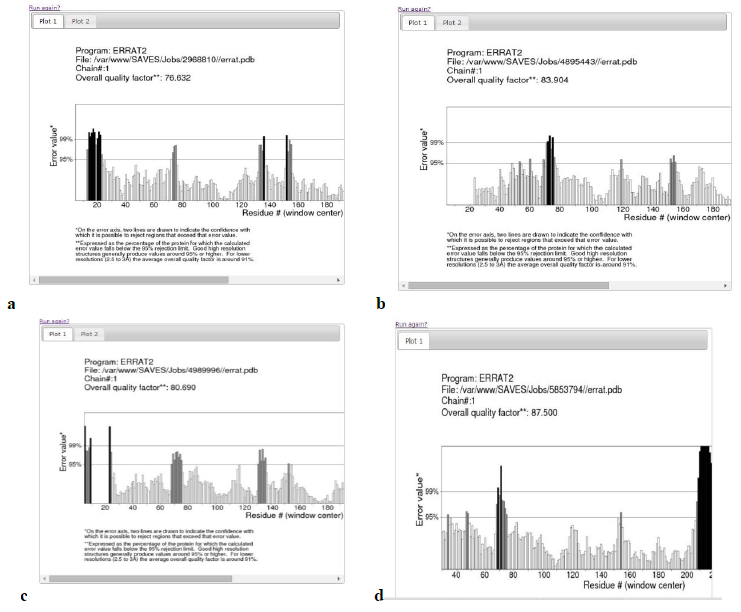
Errat plot- The Errat plots of HLAs are illustrated below. The overall quality factor and RMSD of homology models are shown which further increases the confidence of accepting the homology models.
Table 2: Validation of homology models
|
3D Structures |
Favoured Region |
Allowed Region |
Outlier Region |
Errat Score |
RMSD |
|
Temp: 1XR8 |
369 (97.6%) |
7 (1.9%) |
2 (0.5%) |
94.429 |
0.769 |
|
Query: HLA-B*15:03 |
353 (98.1%) |
7 (1.9%) |
0 (0.0%) |
80.690 |
|
|
Temp: 2XPG |
362 (96.5%) |
13 (3.5%) |
0 (0.0%) |
90.196 |
0.746 |
|
Query: HLA-A*30:02 |
356 (98.1%) |
7 (1.9%) |
0 (0.0%) |
83.904 |
|
|
Temp: 3I6L |
533 (97.4%) |
13 (2.4%) |
1 (0.2%) |
93.220 |
0.444 |
|
Query: HLA-A*23:01 |
355 (97.8%) |
8 (2.2%) |
0 (0.0%) |
76.632 |
|
|
Temp: 4NT6 |
533 (97.4%) |
13 (2.4%) |
1 (0.2%) |
87.989 |
0.566 |
|
Query: HLA-Cw*07:02 |
353 (97.0%) |
10 (2.7%) |
1 (0.3%) |
65.563 |
|
|
After multiple template modeling |
|||||
|
Temp: 4NT6 1QQD 1EFX |
533 (97.4%) 344 (91.7%) 640 (83.0%) |
13 (2.4%) 25 (6.7%) 100 (13.0%) |
1 (0.2%) 6 (1.6%) 31 (4.0%) |
87.989 84.270 79.545 |
0.696 0.8 0.530 |
|
Query: HLA-Cw*07:02 |
264 (97.4%) |
7 (2.6%) |
0 (0.0%) |
87.5 |
|
The RMSD values were calculated using Chimera. In the above analysis the Errat plot score of HLA-Cw*07:02 was not satisfactory, initially. Further model refinement was carried out. The homology model was built by multiple template modeling using 3 different templates.
Molecular docking
Docking Validation
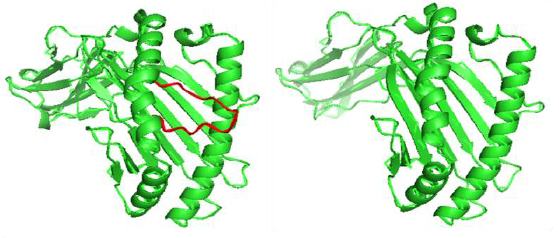
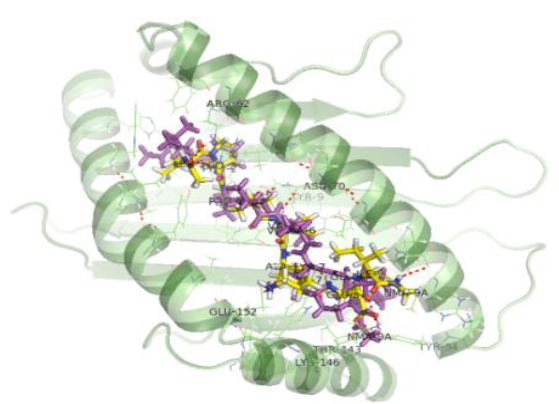
The docking procedure was validated by first removing the co-crystallized epitope available in the PDB structure and redocking it with the same. Docking validation was done using the co crystallized ligand LPPVVAKEI against the HLA-B*51:01 protein. In both cases the peptides bound in the same pose with the HLA and were very much similar conformationally when superimposed.
The interactions shown in the PDB structure are:
H bonds – Tyr 159, Tyr 9, Asn 70 (2 H bonds), Thr 73, Trp 147, Thr 143, Tyr 84, Asn 77, Lys 146
Pi cation –Trp 167
Table 3: Docking validation results
|
Ligand |
MMGBSA DG binding score |
Docking score |
Interactions |
|
LPPVVAKEI |
-158.68 |
-11.59 |
H bonds – Asn 70 (2 H bonds), Tyr 9, Arg 62, Glu 152 (2 H bonds), Thr 143, Lys 146, Glu 76 Salt bridge – Lys 146 (6 similar interactions) |
|
LPPVVAKEI |
-120.54 |
-13.83 |
H bonds – Tyr 159, Asn 70 (2 H bonds), Tyr 9, Thr 73, Tyr 84, Lys 146 (2 H bonds), Trp147 Salt bridge – Lys 146 (10 similar interactions) |
Based on the above results, the peptide docking program of BioLuminate could be validated. The number of interactions which were similar to the co crystallized ligand was maximum in the second case. It has MMGBSA DG binding score -120.54 and therefore this score was taken as a criteria to screen the ligands for all the proteins.
Docking results
- HLA-B*51:01(PDB ID: 1E27) –Following this the 20 ligands were docked against HLA-B*51:01. The top 6 epitopes were selected based on MMGBSA DG binding score, number of interactions with protein that are similar to the experimentally obtained interactions, pose and docking score. 100 conformers were generated for each ligand. The interactions shown in the PDB structure are:
H bonds – Tyr 159, Tyr 9, Asn 70 (2 H bonds), Thr 73, Trp 147, Thr 143, Tyr 84, Asn 77, Lys 146
Pi cation –Trp 167
Salt bridge – Lys 146
Table 4: Best binding scores of different epitopes with HLA-B*51:01
|
Ligands |
MMGBSA DG binding score |
Docking score |
Interactions |
|
FQRTFSIPL |
-152.04 |
-15.78 |
H bonds – Tyr 9, Asn 70 (2 H bonds), Tyr 159, Tyr 99, Tyr 116, Glu 152 (2 H bonds), Trp 147, Asn 77,Tyr 84, Lys 146 |
|
RTFSIPLGV |
-145.75 |
-12.07 |
H bonds – Tyr 159, Tyr 9, Asn 70 (2 H bonds), Trp 147, Thr 73, Glu 152 (2 H bonds) π-π interaction –Thr 73 |
|
TSFFLWVII |
-132.11 |
-10.36 |
H bonds – Tyr 9, Lys 146, Tyr 159, Thr 73, Asn 70 (2 H bonds) π-π interaction – Tyr 159 Pi cation interaction –Arg 62 |
|
STHNTPVYK |
-126.12 |
-10.48 |
H bonds – Glu 152, Tyr 116, Tyr 159, Tyr 7, Tyr 9, Asn 70 (2 H bonds) π-π interaction – Tyr 159 π-cation interaction – Tyr 159 |
|
VIALFCICK |
-123.61 |
-10.74 |
H bonds – Tyr 9, Asn 70 (2 H bond), Tyr 159, Trp 147, Lys 146 (2-H bonds), Glu 152 |
- HLA-B*53:01 (PDB ID: 1A1M)
The interactions shown in the PDB structure before docking includes the following:
H bonds – Asp 114, Tyr 99, Asn 63, Tyr 7, Tyr 171, Arg 62, Asn 77 (2 H bonds), Thr 143, Tyr 84, Trp 147, Lys 146
π-π interactions –Arg 97, Tyr 99
Salt bridge – Lys146
Table 5: Best binding scores of different epitopes with HLA-B*53:01
|
Ligand |
MMGBSA DG binding score |
Docking score |
Interactions |
|
NQDGLICGL |
-124.28 |
-10.94 |
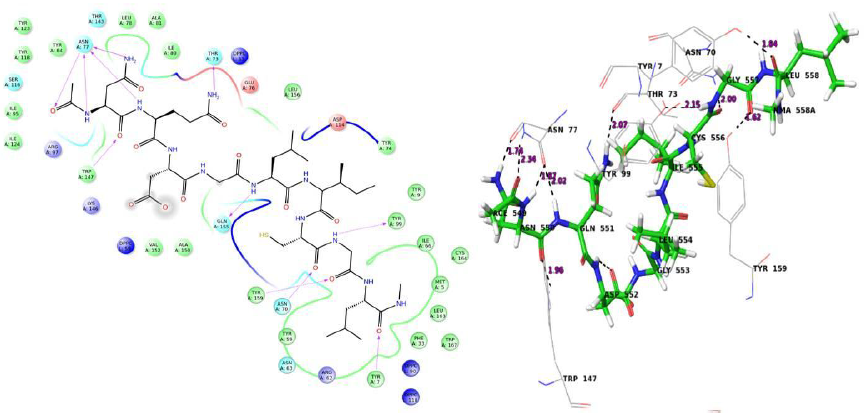
H bonds – Tyr 159, Arg 62, Lys 146, Trp 147, Asn 77(3H bonds), Glu 76, Thr 143 Salt bridge – Lys 146 |
|
RTFSIPLGV |
-120.93 |
-11.60 |
H bonds - Tyr 7, Lys 146, Arg 62, Thr 73, Asn 70, Trp 147, Asn 77, Glu 76 (2 H bonds) |
|
FLWVIILFQ |
-135.89 |
-9.01 |
H bonds – Tyr 84, Thr 143, Trp 147, Lys 146, Tyr 99 (2 H bonds), Arg 62, Gln 155 π-π interaction – Arg 62 |
|
RTSFFLWVI |
-122.25 |
-10.64 |
H bonds -Lys 146, Asn 70, Asn 77, Glu 76, Trp 147 π-π interaction – Arg 97, Tyr 159, Tyr 99 |
|
GYYSTTIRY |
-114.73 |
-12.04 |
H bonds - Leu 163, Arg 62, Asn 70 (2 H bonds), Tyr 74, Asn 77, Arg 97, Trp 147 (2 H bonds), Glu 76 π-π interaction – Tyr 74, Trp 147, Arg 97 |
The interactions shown in the PDB structure before docking includes the following:
H bonds – Ser 77, Asn 80, Lys 146, Thr 143, Tyr 84, Trp 147 (2 H bonds), Gln 155, Tyr 159, Tyr 99, Tyr 171, Tyr 7
Table 6: Best binding scores of different epitopes with HLA-B*35:01
|
Ligands |
MMGBSA DG binding score |
Docking score |
Interactions |
|
NQDGLICGL |
-105.65 |
-13.33 |
H bonds - Tyr 99, Tyr 84, Thr 143, Glu 76, Asn 80 (3 H bonds), Lys 146 (3 H bonds) |
|
STHNTPVYK |
-126.82 |
-10.11 |
H bonds – Tyr 99, Tyr 7, Tyr 171, Arg 97, Ser 77, Asn 70 (2 H bonds), Tyr 9, Lys 146 (2 H bonds), Asn 80 (2 H bonds), Glu 76 (2 H bonds) |
|
RTFSIPLGV |
-122.59 |
-8.92 |
H bonds – Glu 76, Lys 146, Thr 73, Tyr 9, Asn 70, Tyr 99, Gln 155 π-π interaction – Arg 97 |
|
YYSTTIRYQ |
-139.17 |
-10.46 |
H bonds – Gln 155, Arg 62, Tyr 171, Tyr 7, Tyr 99, Arg 97, Lys 146, Ser 77 π-cation interaction – Lys 146 |
|
RTSFFLWVI |
-127.69 |
-11.42 |
H bonds – Lys 146, Asn 80, Glu 76, Trp 147, Asn 70 (2 H bonds), Arg 97, Tyr 9, Tyr 159, Arg 62 |
- HLA-A*02:01 (PDB ID: 3MRE)
The interactions shown in the PDB structure before docking includes the following:
H bonds – Trp 147, Lys 146, Tyr 84, Thr 143, Asp 77, Tyr 99, Lys 66, Glu 63, Tyr 171, Tyr 7
π-cation interaction – Trp 167
Salt bridge – Lys 146, Glu 63
Table 7: Best binding scores of different epitopes with HLA-A*02:01
|
Ligands |
MMGBSA DG binding score |
Docking score |
Interactions |
|
RTSFFLWVI |
-112.60 |
-10.55 |
H bonds - Lys 146, Tyr 116 (2 H bonds), Asp 77, Thr 73 (2 H bonds), Arg 65, Arg 97, Glu 58 π-π interaction – Lys 66 π-cation interaction – Trp 167 |
|
VIALFCICK |
-109.69 |
-9.146 |
H bonds – Lys 146, Lys 146, Arg 97, Tyr99, Tyr 171, Thr 73 π-π interaction – Trp 147 Salt bridge – Glu 63 π-cation interaction – Trp 167 |
|
GYYSTTIRY |
-93.18 |
-9.05 |
H bonds – Lys 146, Tyr 84, Thr 143, Asp 77, Trp 147, Thr 73, Tyr7 (2 H bonds), Trp 167, Thr 163, Arg 65 π-π interaction – Trp 147 π-cation interaction – Trp 167, Lys 66 Salt bridge – Glu 63 |
|
FLWVIILFQ |
-101.15 |
-8.30 |
H bonds – Lys 146, Tyr 84,Asp 77, Trp 147, Lys 66, Trp 167 π-π interaction – Arg 65, HID 70 π-cation interaction – Arg 65 |
|
RTFSIPLGV |
-105.91 |
-7.803 |
H bonds - Lys 146, Trp 147, Glu 58, Thrb73,Tyr 116 (2 H bonds), Arg 97, Arg 65 (2 H bonds), Asp 77 |
- HLA-A*30:02
Table 8: Best binding scores of different epitopes with HLA-A*30:02
|
Ligands |
MMGBSA DG binding score |
Docking score |
Interactions |
|
FLWVIILFQ |
-109.60 |
-8.52 |
H bonds – Thr 167, Tyr 108, Lys 170, Trp 171, Asn 90, Gln 86 π-π interaction – Trp 191, Tyr 183 |
|
AEGVVAFLI |
-100.73 |
-10.48 |
H bonds – Trp 171, Asp 101 (2 H bond) Lys 170 (2 H bonds), Gln 179, Glu 87 π-π interaction – Tyr 123, Tyr 183 Salt bridge – Lys 170 |
|
FQRTFSIPL |
-81.60 |
-8.99 |
H bonds – Arg 89, Glu 87, Gln 179, Thr 187, Asp 101 π-π interaction – Tyr 183 |
|
TFAEGVVAL |
-73.03 |
-8.44 |
H bonds – Asp 101, Lys 170, Tyr 123, Tyr 183, Glu 87, Trp 176, Trp 171 π-cation interaction – Lys 170 |
|
SFFLWVIIL |
-62.58 |
-9.449 |
H bonds – Thr 104, Lys 170, Asn90,Gln 86,Trp 191 π-π interaction – Trp 176 |
- HLA-A*23:01
Table 9: Best binding scores of different epitopes with HLA-A*23:01
|
Ligands |
MMGBSA DG binding score |
Docking score |
Interactions |
|
RTSFFLWVI |
-96.97 |
-6.51 |
H bonds – Thr 187, Asp 190 (2 H bonds), Gln 179, Thr 97 |
|
FQRTFSIPL |
-96.34 |
-10.51 |
H bonds – Glu 86 (4 H bonds), Lys 90 (2 H bonds), Tyr 195, Tyr 183, Lys 170 Salt bridge – Glu 86 |
|
FFLWVIILF |
-92.53 |
-8.81 |
H bonds – Thr 97, Thr 187, Tyr 195 |
|
RTFSIPLGV |
-89.16 |
-6.45 |
H bonds – Asp 190, Tyr 195, Trp 171 |
|
YYSTTIRYQ |
-84.59 |
-9.88 |
H bonds – Glu 87 (2 H bonds), Thr 187, Gln 179, Asn 101,Lys90 |
- HLA-B*15:03
Table 10: Best binding scores of different epitopes with HLA-B*15:03
|
Ligands |
MMGBSA DG binding score |
Docking score |
Interactions |
|
RTSFFLWVI |
-101.69 |
-6.89 |
H bonds –Trp 180, Ser 101, Trp 191, Glu 82 (2 H bonds) π-π interaction – Trp 180 (2 bonds), Tyr 183 |
|
TTIGEWAFW |
-81.22 |
-8.08 |
H bonds – Lys 170, Glu 82, Ser 101, Glu 176, Trp 171, Thr 97, Gln 89 |
|
FQRTFSIPL |
-105.38 |
-7.16 |
H bonds – Arg 86, Glu 176 (2 H bonds), Gln 179 (2 H bonds), Trp 171, Lys 170 |
|
GYYSTTIRY |
-105.33 |
-7.72 |
H bonds – Gln 179, Glu 176 (2 H bonds), Asn 94, Tyr 23, Tyr 108, Thr 167 π-π interaction –Trp 171 Salt bridge – Glu 176 |
|
RTFSIPLGV |
-89.77 |
-5.95 |
H bonds – Glu 82,Arg 86, Glu 176, Ser 101 π-cation interaction – Tyr 98, Arg 121 |
- HLA-Cw*07:02
Table 11: Best binding scores of different epitopes with HLA-Cw*07:02
|
Ligands |
MMGBSA DG binding score |
Docking score |
Interactions |
|
RTFSIPLGV |
-130.20 |
-8.10 |
H bonds – Asp 33 (2 H bonds), Arg 121, Gln 94 (4 H bonds), Asn 104, Lys 90, Glu 87 Salt bridge – Asp 33 |
|
FQRTFSIPL |
-127.58 |
-10.55 |
H bonds – Asn 104, Asp 138, Gln 94, Asp 33 (2 H bonds), Arg 93, Arg 121 π-π interaction – Arg 86 π-cation interaction – Arg 86 |
|
FAFHKEGAF |
-120.91 |
-8.41 |
H bonds – Asp 98, Gln 179, Tyr 31, Glu 87, Lys 90, Gln 94 π-π interaction –Tyr 31, Tyr 183 π-cation interaction – Arg 121, Arg 93 |
|
FLWVIILFQ |
-118.97 |
-11.64 |
H bonds – Tyr 31, Glu 87, Gln 94 (2 H bonds), Tyr 108, Lys 90, Lys 170 π-π interaction – Tyr 91 π-cation interaction – Lys 170 |
|
YYSTTIRYQ |
-109.45 |
-11.50 |
H bonds – Glu 87, Tyr 31, Asp 138, Asp 33, Gln 179 (2 H bonds), Arg 93 (2 H bonds), Tyr 108, Thr 167, Asp 98, Lys 90 π-π interaction – Tyr 91, Tyr 183 |
Apart from the above epitopes, TFAEGVVAF, NQDGLICGL, GYYSTTIRY, VIALFCICK, STHNTPVYK and GTNETEYLF were also found to have good pose, interaction and good score when docked against HLA-Cw*07:02. The cut off for MMGBSA DG binding score was considered as -120 and hence these epitopes were screened off.
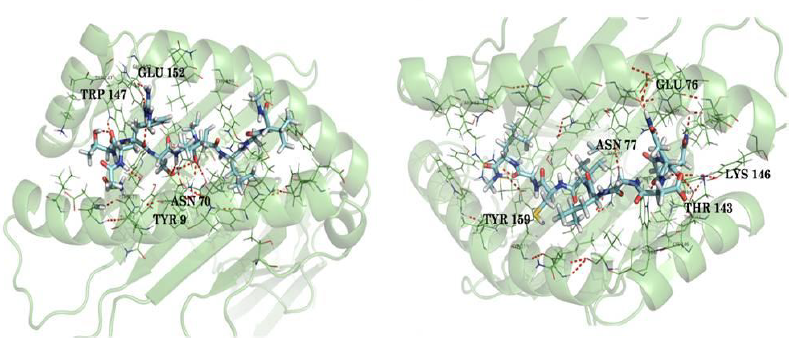
In docking analysis, some common H bond interactions were found, mainly with Tyr 9, Asn 70, Lys 146, Tyr 159, Trp 147 and Asn 77 for different HLA proteins. Salt bridge was found in many cases with Lys 146. The common interactions were found to be less in case of proteins that were modeled. The peptides were capped with N-methyl amide and acetyl groups. The residues in bold indicates the bond with these caps. From the analysis it can be concluded that FQRTFSIPL, RTFSIPLGV, NQDGLICGL, STHNTPVYK, FLWVIILFQ, VIALFCICK and RTSFFLWVI can be possible epitopes that can trigger a T cell immune response in humans. Molecular dynamics was carried out to validate docking results.
Molecular dynamics
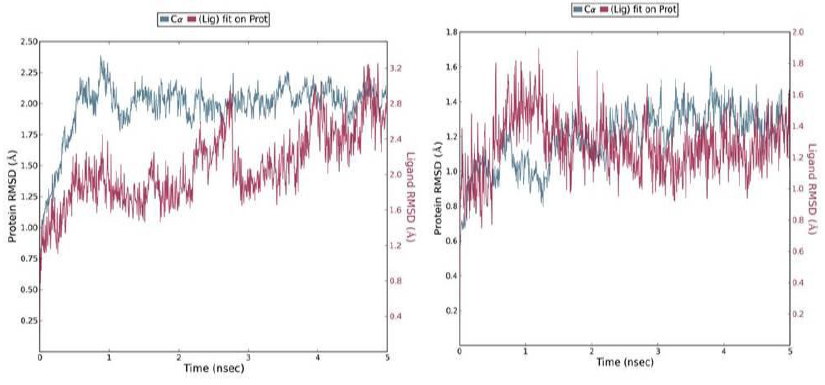
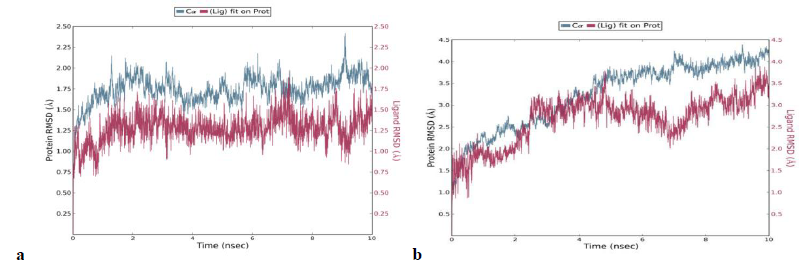
A 5 ns simulation of the docked complex structure of HLA-B*53:01 with ligand NQDGLICGL was performed to obtain a dynamical picture of the conformational changes that occur in aqueous solutions. The simulation was further extended to 10ns under the same conditions. The RMSDs of the trajectory of protein and ligand is depicted below.
The RMSD showed that initially there was a greater fluctuation but after 7ns, the simulation converges. It indicates that the simulation has almost equilibrated throughout 10ns.
RMSD of the PDB structure 1A1M (HLA-B*53:01) with co crystallized ligand TPYDINMQML was carried out for 10ns. The RMSD showed an equilibrated simulation as depicted below.
Further, the RMSD of ligand (RTFSIPLGV) with HLA-Cw*07:02 was analyzed and the graph obtained is shown below. The RMSD was found to be highly fluctuating, hence the result was not found to be satisfactory in this case.
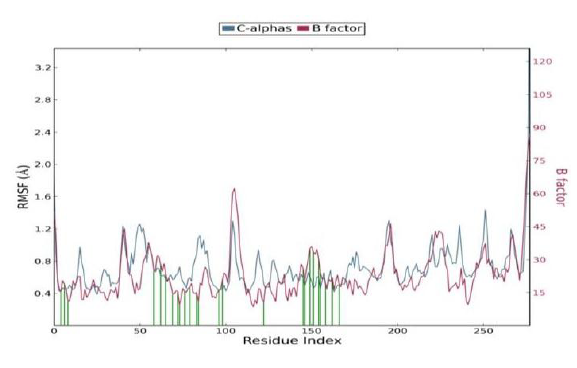
Protein RMSF
The Root Mean Square Fluctuation (RMSF) is useful for characterizing local changes along the protein chain. Protein residues that interact with the ligand are marked with green-colored vertical bars.
In this plot, peaks indicate areas of the protein that fluctuate the most during the simulation. It is observed that the tails (N- and C-terminal) fluctuate more than any other part of the protein. Secondary structure elements like alpha helices and beta strands are usually more rigid than the unstructured part of the protein, and thus fluctuate less than the loop regions. The RMSF is correlated with the PDB B factor. Due to the difference between the RMSF and B factor definitions, one-to-one correspondence should not be expected. However, the simulation results should parallel the crystallographic data as shown below.
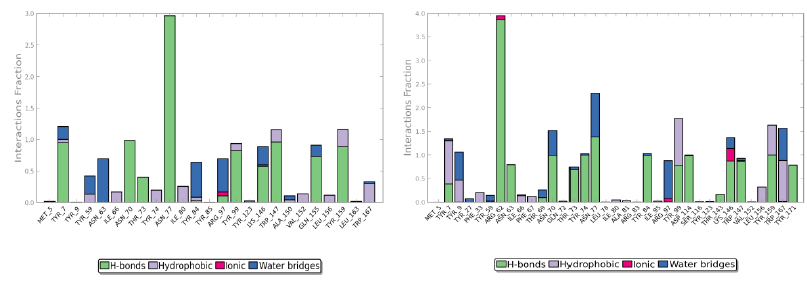
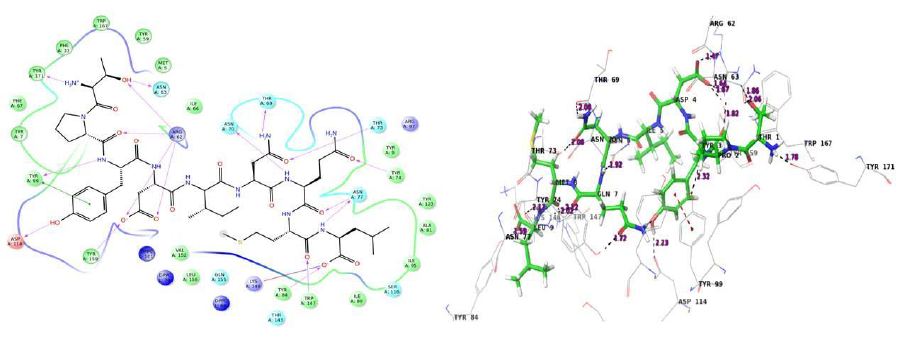
Protein interactions with the ligand can be monitored throughout the simulation. These interactions can be categorized by type and summarized, as shown in the plot above. Protein-ligand interactions (or 'contacts') are categorized into four types: hydrogen bonds, hydrophobic (π-cation and π-π interaction), ionic and water bridges.
The value 0.5 indicates 50% of the simulation time the specific interaction was maintained. In the above graph Asn 77 shows approximately 3.0 interactions fraction, as it makes 3 H bond with the ligand residues. Tyr 7, Tyr 99, Trp 147, Asn 70, Tyr 159 and Gln 155 are other residues.
The Arg 62 residue of the PDB structure forms about 4 H bonds with the co crystallized ligand. In the docked structure even though this residue is not making any significant interactions, the ligand makes good contacts with other important residues of the protein.
Conclusion
Development of effective therapeutics for Ebola virus remains a high priority in the present scenario. This study is based on epitope focused vaccine design against Ebola virus. The study focused mainly on T-cell epitopes as the algorithms for identifying B-cell epitopes are not highly reliable. Different individuals respond to a disease differently and this is based on the HLA alleles present in them, which differs in each person.
The major task was to find the suitable epitopes. It involved screening through different online servers. At this stage, it was necessary to specify HLA alleles as epitope-HLA interaction differs with different alleles.
Eight HLA alleles were selected, based on its occurrence in African and World population. HLA-B*53:01, HLA-B*51:01, HLA-B*35:01, HLA-A*02:01 are those alleles whose PDB structures were available and HLA-A*30:02, HLA-A*23:01, HLA-B*15:03 and HLA-Cw*07:02 needed to be modeled using Modeler 9.14.
The selected top 20 epitopes were docked with each HLA and FQRTFSIPL, RTFSIPLGV, NQDGLICGL, STHNTPVYK, FLWVIILFQ, VIALFCICK and RTSFFLWVI were found to have good pose, interaction and high MMGBSA binding score. Hence, they can be considered as suitable vaccine candidates. Further, molecular dynamics studies were carried out using Desmond, for the docked complex HLA-B*53:01-NQDGLICGL. The interactions found were satisfactory and the complex was equilibrated during the simulation through 10ns. So, it can be a suitable vaccine candidate. At the same time the interactions between HLA-Cw*07:02 and RTFSIPLGV was found to be lesser stable. As seen in the history of reverse vaccinology, multiple epitope based vaccines show improved results. The addition of certain adjuvants also increase the efficacy of vaccines. The interactions may be increased by any of these methods. Further invitro studies are needed. Finally, it can be concluded that FQRTFSIPL, RTFSIPLGV, NQDGLICGL, STHNTPVYK, FLWVIILFQ, VIALFCICK and RTSFFLWVI are promising epitope vaccine candidates.
Acknowledgment
We would like to acknowledge Director, NIPER, Mohali for providing the facility for carrying out the work. The financial assistance provided by the Department of Science and Technology (DST), New Delhi, and The Council of Scientific & Industrial Research (CSIR), New Delhi is duly acknowledged.
References
- Centers for Disease Control and Prevention. Bioterrorism agents/diseases. http://www.bt.cdc.gov/agent/agentlist-category.asp#a (accessed March 20, 2016).
- Feldmann H, Geisbert TW. Ebola haemorrhagic fever. The Lancet 377 (2011): 849-862.
- Lozano R, Naghavi M, Foreman K, et al. Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: a systematic analysis for the Global Burden of Disease Study 2010. The Lancet 380 (2012): 2095-2128.
- Murray CJ, Vos T, Lozano R, et al. Disability-adjusted life years (DALYs) for 291 diseases and injuries in 21 regions, 1990–2010: a systematic analysis for the Global Burden of Disease Study 2010. The Lancet 380 (2012): 2197-2223.
- Zentner G. Building a Better Vaccine. Science Spotlight 4 (2014).
- Correia BE, Bates JT, Loomis RJ, et al. Proof of principle for epitope-focused vaccine design. Nature 507 (2014): 201-206.
- "Partnership for Research on Ebola Vaccines in Liberia (PREVAIL)". 24 June 2015. Retrieved 10 September 2015. https://clinicaltrials.gov/ct2/show/NCT02344407
- Marzi A, Feldmann H, Geisbert TW, et al. Vesicular Stomatitis Virus-Based Vaccines for Prophylaxis and Treatment of Filovirus Infections. Journal of Bioterrorism & Biodefense 1 (2011): 2157-2526.
- World Health Organization Ebola vaccines, therapies and diagnostics. Available at: http://www.who.int/medicines/emp_ebola_q_as/en. Accessed June 10, 2015
- "A Safety and Immunogenicity Study of Heterologous Prime-Boost Ebola Vaccine Regimens in Healthy Participants". 23 July 2015. Retrieved 15 October 2015. https://clinicaltrials.gov/ct2/show/NCT02313077
- Pizza M, Scarlato V, Masignani V, et al. Identification of vaccine candidates against serogroup B meningococcus by whole-genome sequencing. Science 287 (2000): 1816-1820.
- Kelly DF, Rappuoli R. Reverse vaccinology and vaccines for serogroup B Neisseria meningitidis. InHot Topics in Infection and Immunity in Children II (2005): 217-223. Springer, Boston, MA.
- Santos A, Ali A, Barbosa E, et al. The reverse vaccinology-A contextual overview. IIOABJ 2 (2011): 8-15.
- Bowman BN, McAdam PR, Vivona S, et al. Improving reverse vaccinology with a machine learning approach. Vaccine 29 (2011): 8156-8164.
- Kanampalliwar AM, Soni R, Girdhar A, et al. Reverse Vaccinology: Basics and Applications. J Vaccines Vaccin 4 (2013): 1-5.
- Ferenczy MW. Prophylactic vaccine strategies and the potential of therapeutic vaccines against herpes simplex virus. Current Pharmaceutical Design 13 (2007): 1975-1988.
- Doytchinova IA, Flower DR. VaxiJen: a server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinformatics 8 (2007): 1-7.
- Huang J, Honda W. CED: a conformational epitope database. BMC Immunology 7 (2006): 1-8.
- Potocnakova L, Bhide M, Pulzova LB. An introduction to B-cell epitope mapping and in silico epitope prediction. Journal of Immunology Research 2016 (2016).
- Alberts B, Johnson A, Lewis J, et al. T cells and MHC proteins. InMolecular Biology of the Cell. 4th edition (2002).
- De Groot AS. Immunomics: discovering new targets for vaccines and therapeutics. Drug Discovery Today 11 (2006): 203-209.
- De Groot AS, Berzofsky JA. From genome to vaccine--new immunoinformatics tools for vaccine design. Methods (San Diego, Calif.) 34 (2004): 425-428.
- Van Regenmortel MH. Immunoinformatics may lead to a reappraisal of the nature of B cell epitopes and of the feasibility of synthetic peptide vaccines. Journal of Molecular Recognition: JMR 19 (2006): 183-187.
- L Dudek N, Perlmutter P, Aguilar I, et al. Epitope discovery and their use in peptide based vaccines. Current Pharmaceutical Design 16 (2010): 3149-3157.
- Agadjanyan MG, Ghochikyan A, Petrushina I, et al. Prototype Alzheimer’s disease vaccine using the immunodominant B cell epitope from β-amyloid and promiscuous T cell epitope pan HLA DR-binding peptide. The Journal of Immunology 174 (2005): 1580-1586.
- Rammensee HG, Bachmann J, Emmerich NP, et al. SYFPEITHI: database for MHC ligands and peptide motifs. Immunogenetics 50 (1999): 213-219.
- Schönbach C, Koh JL, Flower DR, et al. An update on the functional molecular immunology (FIMM) database. Applied Bioinformatics 4 (2005): 25-31.
- Bhasin M, Singh H, Raghava GP. MHCBN: a comprehensive database of MHC binding and non-binding peptides. Bioinformatics 19 (2003): 665-666.
- Reche PA, Zhang H, Glutting JP, et al. EPIMHC: a curated database of MHC-binding peptides for customized computational vaccinology. Bioinformatics 21 (2005): 2140-2141.
- Saha S, Bhasin M, Raghava GP. Bcipep: a database of B-cell epitopes. BMC Genomics 6 (2005): 1-7.
- Schlessinger A, Ofran Y, Yachdav G, et al. Epitome: database of structure-inferred antigenic epitopes. Nucleic Acids Research 34 (2006): D777-D780.
- Toseland CP, Clayton DJ, McSparron H, et al. AntiJen: a quantitative immunology database integrating functional, thermodynamic, kinetic, biophysical, and cellular data. Immunome Research 1 (2005): 1-2.
- Doytchinova IA, Flower DR. VaxiJen: a server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinformatics 8 (2007): 1-7.
- Larsen MV, Lundegaard C, Lamberth K, et al. Large-scale validation of methods for cytotoxic T-lymphocyte epitope prediction. BMC Bioinformatics 8 (2007): 424.
- Rammensee HG, Bachmann J, Emmerich NP, et al. SYFPEITHI: database for MHC ligands and peptide motifs. Immunogenetics 50 (1999): 213-219.
- Dimitrov I, Flower DR, Doytchinova I. AllerTOP-a server for in silico prediction of allergens. InBMC Bioinformatics 14 (2013): 1-9.
- Tubert-Brohman I, Sherman W, Repasky M, et al. Improved docking of polypeptides with Glide. Journal of Chemical Information and Modeling 53 (2013): 1689-1699



 Impact Factor: * 3.1
Impact Factor: * 3.1 CiteScore: 2.9
CiteScore: 2.9  Acceptance Rate: 11.01%
Acceptance Rate: 11.01%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 
