Interleukin-6 Gene Promoter Region Polymorphism in Patients with Dilated Cardiomyopathy
Article Information
Sumbal Arooj1,2*, Muhammad Khan3, Ali Raza4
1Department of Biochemistry, Qauid-i-Azam University Islamabad, Pakistan
2Department of Biochemistry, University of Sialkot, Pakistan.
3Department of Oncology, the First Affiliated Hospital, Anhui Medical University, Hefei, Anhui Province 230022, People’s Republic of China
4Department Oo Paediatrics, Mayo Hosoital, King Edward Medical University, Lahore
*Corresponding author: Sumbal Arooj, Department of Biochemistry, Qauid-i-Azam University Islamabad, Pakistan and Department of Biochemistry, University of Sialkot, Pakistan
Received: 31 July 2019; Accepted: 16 August 2019; Published: 19 August 2019
Citation: Sumbal Arooj, Muhammad Khan, Ali Raza. Interleukin-6 Gene Promoter Region Polymorphism in Patients with Dilated Cardiomyopathy. Cardiology and Cardiovascular Medicine 3 (2019): 329-339.
View / Download Pdf Share at FacebookAbstract
Background: Dilated cardiomyopathy (DCM) is a primary myocardial disease characterized by left ventricular or biventricular dilatation and impaired myocardial contractility. IL-6 is a pro-inflammatory cytokine that has been found to play an important role in the pathogenesis of DCM. The present study endeavors to explore the association of IL-6 -174 G/C polymorphism in patients with dilated cardiomyopathy, in addition to several other contributing risk factors in Pakistani population.
Method: The study population comprised of 76 patients (54 males and 22 Females) and 66 controls (47 males and 19 Females). Biochemical characteristics were analyzed to detect serum lipid profile. Genotyping was carried out by polymerase chain reaction–restriction fragment length polymorphism (PCR-RFLP). The data was analyzed for statistical significance using student t-test, chi-square analysis.
Result: Comparison of wild type (GG) and (GC+CC) variant for IL-6 -174 G/C polymorphism showed that GC+CC was more common in patients’ group (20%) as compared to controls (4.5%). Similarly, significant association of variant allele C with DCM patients was found (OR = 4.708, 95% CI = 1.331-16.648, P = 0.0175). BMI was significantly higher among patients as compared to controls (P = 0.0004). The lipid profile of studied population showed significantly increased serum cholesterol (P < 0.0001), LDL-C levels (P < 0.0001) and HDL-C (P < 0.0001) in DCM patients as compared to controls. However, no significant difference was observed for TG and vLDL-C between patients and controls.
Conclusion: Our findings indicated that IL-6 -174 G/C gene polymorphism may be associated with development of DCM.
Keywords
Dilated Cardiomyopathy; Interleukine-6; Gene polymorphism
Article Details
1. Introduction
Cardiomyopathy is disease of heart muscle involving three major abnormal findings such as chamber size, wall thickness, and functional contractility [1]. DCM is one of such abnormality characterized by dilation and impaired contractility of left or both ventricles, along with progression in functional and structural changes of systolic dysfunction resulting in progressive heart failure with sudden or heart failure?related death as final outcome [2]. It is more common in men than in women [3]. A number of genetic and environmental factors are involved in induction of myocardial damage associated with DCM and is manifested clinically in third or fourth decade of life [3,4]. Several years of research on the etiology and development of DCM has failed to identify the cause of DCM. Nonetheless, familial and genetic disorders, infectious and toxicity related processes, autoimmunity, and inflammation have been associated with the development of DCM. In most cases, the causal mechanism of disease is poorly understood, and hence the term idiopathic has been used with DCM as “Idiopathic DCM (IDCM)” [4].
Increasing evidence suggests several inflammation or immune response associated cytokines are participating in the pathogenesis of DCM and even congestive heart failure [5, 6]. Moreover, gene polymorphism of some of these proinflammatory cytokines such IL-6, IL-31 and TNF-a are associated with worst prognosis or disease severity of DCM or heart failure [5-11]. The interleukin 6 (IL-6) gene in humans is located in the short arm of chromosome 7 and two functional polymorphisms in the IL-6 promoter have been identified - one at position-174 and one at position-572 (-174G/C and -572G/C). IL-6-174G/C has been associated with DCM, CHD and blood pressure risk. The carriers of C allele were associated with increased risk for DCM, CHD risk and blood pressure [7,8,10]. Interleukin-6 gene polymorphisms have also been associated with other diseases such as systemic-onset juvenile chronic arthritis, sporadic Alzheimer's disease and rheumatoid arthritis [12-14]. Thus, IL-6 gene polymorphism plays an important role in the pathogenesis of various diseases including DCM. Our study aims to identify the association between IL-6-174G/C and DCM in Pakistani population. In addition, several contributing risk factors associated with DCM would also be identified.
2. Methodology
2.1 Study Population
This study was comprised of 76 DCM patients and 66 healthy controls. The criterions were established to collect the blood samples of patients and controls. The patients were recruited from the cardiology outpatient departments of multiple cardiology centers in Rawalpindi/Islamabad, Pakistan namely Federal Government Poly Clinic (Post Graduate Medical Institute) Islamabad; Armed Forces Institute of Cardiology/National Institute of Heart Diseases, Rawalpindi; and Pakistan Institute of Medical Sciences, Islamabad). Diagnosis for DCM patients was made on the basis of history and physical examination, chest X-ray demonstration of generalized ventricular enlargement, electrocardiogram (ECG) showing non-specific ST segment and T-wave changes, and echocardiographic evidence of left ventricular enlargement, left ventricular systolic dysfunction (ejection fraction ≤ 40%) and end-diastolic diameter ( >34 mm/m2), in accordance to World Health Organization Guidelines [15-18].
Exclusion criteria included patients diagnosed with systemic arterial hypertension (>160/100 mmHg), coronary heart disease, systemic diseases, pericardial diseases, congenital heart disease, Cor-pulmonale and clinical, sustained and rapid supraventricular arrhythmias. Clinically healthy individuals (number [n]=66; mean [SD] age: 54.7 [13.4] years) with normal ECG and echocardiography, showing no symptoms of any concomitant disease, and having no familial history of cardiomyopathy or other heart ailments, were selected as control subjects. Recruitment was done from regions of patients’ areas and unrelated healthy individuals were selected keeping in view the genetic substructure in the Pakistani population. This study was reviewed and approved by the Institutional Review Board, Quaid-i-Azam University, Islamabad. Informed consent was obtained from all participants of the study in accordance with the Declaration of Helsinki of 1975, 1997 revision [19].
2.2 Blood Sample Collection
Samples (10 ml peripheral venous blood) were collected from each patient and control with a sterilized syringe using aseptic vein puncture technique. After collection, half of the volume of blood (5 ml) was immediately transferred to the standard potassium EDTA anticoagulant tube. After proper labeling, these tubes were transferred to dry ice box (4 0C) for transportation to our laboratory. These samples were used for DNA extraction and genetic studies. The remaining half volume (5 ml) was placed in plain (autoclaved and sterilized) serum tubes and allowed to clot for some time. The samples were centrifuged at 4000 rpm for 5 min to separate serum. Serum was separated in eppendrof tubes (0.5ml aliquots). These aliquots were transferred to low temperature storage (-20 0C). Serum samples were used for biochemical analysis.
2.3 Biochemical Analyses
AMP Diagnostics kits (AMEDA Laboradignostik GmbH, Graz, Austria) was used for biochemical analyses for the determination of total cholesterol (TC), triglycerides (TG), low-density cholesterol (LDL-C), and high-density cholesterol (HDL-C) from serum samples using Vitalab Selectra E Chemistry Analyzer (Vitalab, Hoogerheide, the Netherlands). The assays were performed according to the manufacturer’s instructions using standard enzymatic techniques.
2.4 Extraction of DNA and Genotyping of -174 G/C IL-6 SNP
IL-6 gene polymorphism was detected by PCR using the following primers of 20 bp size. Total amplified region was of 408 base pairs
F: 5’ GCGATGGAGTCAGAGGAAAC 3’
R: 5’ ATCTTTGTTGGAGGGTGAGG 3’
PCR was performed in 0.2 ml PCR tubes (Axygen, USA) containing 50 µL total reaction mixture. The reaction mixture was prepared by adding 3 µL of DNA sample, 5 µL of 10 X buffer (200 mM (NH4)2SO4, 750 mM of Tris-HCL pH 8.8 and 0.1 % Tween 20), 4 µl MgCl2 (25 mM), 1 µl dNTPs (10 mM), 2.5 µl of each forward and reverse primer (0.1 µM), 0.5 µl Taq DNA polymerase (5 units/µl, Fermentas, UK) and 31.5 µl of PCR water. For amplification purpose, high throughput thermal cycler (Applied Biosystems GeneAmp® PCR System 9700) (Life Technologies, USA) was used.
Amplified DNA products were analyzed by 2% agarose gel electrophoresis, which was prepared by dissolving 1.0 g of agarose in 50 ml of 1X TBE buffer. Amplified PCR products were mixed with loading dye (0.15 % bromophenol blue with 40 % sucrose solution). Electrophoresis was performed at 100 volts (50 mA) for 30 minutes by using 1X TBE running buffer. PCR fragments were detected by placing the gel on UV transilluminator (TFX-35 M- Life Technologies, USA).
Genotyping of -174 G/C polymorphism was done using restriction fragment length polymorphism (RFLP) technique. 1 µl (6 U/µl) of NlaIII restriction enzyme mixed with 2 µl of 10X Fast Digest buffer, was added to 10 µl of PCR products. The reaction volume of 20 µl was prepared by adding PCR water. The mixture was then incubated at 37 oC for 16-18 hours.
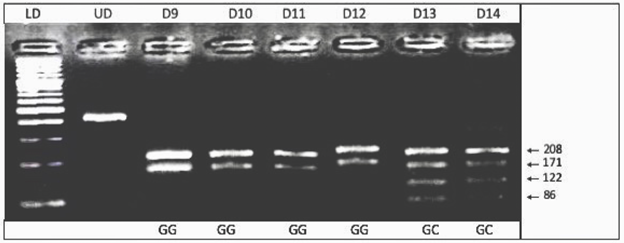
Digested PCR products were analyzed by 3% agarose gel electrophoresis. 3% (w/v) agarose gel was prepared by dissolving 1.5 g of agarose in 50 ml of 1X TBE buffer. The DNA fragments were visualized by the gel documentation system (Syngene, USA) and genotypes were determined. The GG genotype showed 3 bands of 208 bp, 171 bp, and 29 bp; the CC genotype showed bands of 171 bp, 122 bp, 86 bp, and 29 bp; whereas the heterozygous GC showed 208 bp, 171 bp, 122 bp, 86 bp, and 29 bp bands on the gels (Figure 1).
Figure 1: Electropherogram of ethidium bromide stained 3% agarose gel showing genotype pattern obtained with NlaIII restriction digest at -174 G/C IL-6 gene polymorphism. Lane LD 100bp shows DNA ladder. Lane UD represents undigested amplified PCR product of 408bp. The genotypes of patients are shown in bottom legend. The GG genotype showing bands of 208bp and 171bp while GC showing bands of 208bp, 171bp, 122bp and 86bp. Band of 29 cannot be seen.
2.5 Statistical Analysis
All the statistical analyses were carried out using the GraphPad Instat 3.0 (GraphPad Software, Inc., USA). Baseline characteristics of the study population were reported as frequencies and percentages, and the clinical parameters were reported as means with standard deviations. Comparisons of the baseline variables between control and patient subjects were carried out using Fisher’s exact test. To compare the systolic BP, diastolic BP, BMI and the levels of cholesterol, triglyceride, LDL, HDL, and vLDL between the two groups, unpaired student’s t test was performed. From the observed counts, allelic and genotypic frequencies between the two groups were determined by Fisher’s exact test and Chi-square test. The statistical significance between the samples from the study groups was analyzed by the chi-square and Student t tests where indicated. Deviation from the Hardy–Weinberg equilibrium was evaluated by comparing observed and expected genotype frequencies via the chi-square test separately in cases and controls. The genotype and allele frequencies between the patient and control groups were compared by a chi-square test. The criterion for significance was P<0.05.
3. Results
3.1 Baseline Characteristics of Study Population
The patient group contained 22 (29 %) females and 54 (71 %) males while control group contained 19 (29 %) females and 47 (71 %) males in the respective groups (P = 1.0000). Smoking (P = 0.4042) and age (P = 0.8526) showed no significant difference between two groups. Overweight showed significant difference between patients and controls (P = 0.0009).
3.2 Clinical and Biochemical Factors Association
Systolic BP, diastolic BP and body mass index also showed significance between patients and controls (P < 0.05). Mean total cholesterol, LDL-C and HDL-C showed significant differences in patients as compare to controls (P < 0.0001). Other biochemical parameters, triglycerides and vLDL-C, showed no significant variations (P > 0.05, Table 3.3).
|
Genotype |
Controls |
Patient |
P Value |
|
n = 66 |
n = 76 |
||
|
GG |
63 (95.5%) |
61 (80%) |
0.0132 |
|
GC |
3 (4.5%) |
15 (20%) |
|
|
CC |
0 (0%) |
0 (0%) |
Table 3.3: Genotype frequencies of the IL-6 -174 G/C T polymorphism in the study population.
P Value calculated by Chi-square test.
3.3 Genotype Distribution
Tables 3.4 and 3.5 show genotype frequencies of healthy control subjects and DCM patients. Significant difference was observed in the genotype frequency. The frequency of GG genotype among patients and control group was n=61 (80%) and n=63 (95.5%) respectively. Whereas, prevalence of GC genotype in patients was n=15 [20] compared with n=3 (4.5%) in control subjects. The CC homozygous genotype was not present in patient as well as in control groups.
Similarly, allele frequencies of G for patients (90%) as compared to controls (98%) and allele frequencies of C for patients (10%) as compared to controls (2%), showed statistically significant difference between control subjects and DCM patients (P = 0.0175, OR = 4.708, 95% CI = 1.331-16.648).
|
Genotype |
Controls n (%age) |
Patients n (%age) |
OR |
95% CI |
P Value |
|
G C |
129 (98%) 3 (2%) |
137 (90%) 15 (10%) |
4.708 |
1.331-16.648 |
0.0175 |
Table 3.4: Allele frequencies of the IL-6 -174 G/C polymorphism in the study population.
Values are given in numbers
P value calculated by Chi-square test
OR, Odds ratio; CI, Confidence Interval
4. Discussion
Cytokines are produced by various cell types and often show overlapping activities regulating proliferation or differentiation, depending on the type and developmental state of the target cells involved. Cytokines, like interleukin-6 (IL-6), IL-1, and TNFα, are elevated in most inflammatory states and have been termed as pro-inflammatory cytokines. Therefore, these cytokines have also been recognized as targets for therapeutic intervention [20]. DCM associated myocardial dysfunction have also been reported with elevated levels of several of these pro-inflammatory cytokines including IL-1α, IL-1β, IL-6, IL-31, TNF-α, and INF-γ [10,11]. Interleukin-6 (IL-6) and its receptor membrane glycoprotein gp130 have shown to be elevated in IDCM [6,21]. Membrane glycoprotein gp130 act as a common receptor and signal transducer subunit for other members of IL-6 family namely IL-11, leukemia inhibitory factor (LIF), oncostatin M (OSM), ciliary inhibitory factor (CNTF), cardiotropin-1 (CT-1), cardiotrophin-like related cytokine and stimulating neurotrophin-1/B-cell stimulating factor 3 (NNT-1), neuropoietin (NPN), IL-27, with the exception of IL-30 [11,20].
Gene polymorphism of some of these cytokines (IL-6 and TNF-α) have been associated with DCM [10,22,23]. The human IL-6 gene is polymorphic, and one of the characterized polymorphisms is a single base change (G/C) at the promoter site -174 [24]. Several studies have reported IL-16 -174 G/C polymorphism association with a number of diseases including CHF and IDCM. In our study, comparison of wild type (GG) genotype (61%) and variant (GC+CC) genotype (15%) between controls and patients showed statistical significance (P = 0.0132; Table 3.3). Similarly, the allelic frequency of IL-6 -174 gene polymorphism showed higher prevalence of C allele in patients (10%) as compared to controls (2%) and the G allele was more prevalent in controls (98%) as compared to patients (90%) (P = 0.0175; Table 3.4).
The C allele at position-174 in the promoter of the interleukin 6 (IL-6) gene has been associated with reduced gene expression and reduced plasma levels of IL-6 (25). This seems to be in contradiction of the assumption that IL-6 –174 G/C polymorphism may be associated with increased serum IL-6 levels [22,26-33]. Age of the patients, in the studies that revealed increased serum IL-6 levels association with C allele, was at least over 65 [26,27]. Bruunsgaard, et al. showed C allele association with increased serum IL-6 levels but this association was only in current non-smokers, and no association was established with mortality [27]. Panoulas, et al. also reported significantly increased IL-6 levels in C-allele carriers [14.02 (3.21–38.81) vs. 4.48 (2.25–16.5), p = 0.028], and a trend for an interaction between ever smoking and IL6 C-allele carriers on CVD [28]. In presence of pre-existing major disease, C+ carrier status was associated with increased baseline serum IL-6 [odds ratio 2.01; 95%CI: 1.25–3.22], however, C+ carrier status did not affect both serum IL-6 and mortality in the absence pre-existing major disease [29]. C allele carriers (genotype -572C compared to -572GG (P=0.03), and genotype -174CC allele compared to -174G allele carriers (P=0.04) were also reported to have significantly higher peak IL-6 levels 6 hours after CABG while all the genotypes had similar baseline IL-6 levels [30]. Kriplin, et al. showed IL-6 –174 G/C polymorphism association with serum IL-6 levels in neonates but not in adults [31].
Other studies have reported higher serum IL-6 levels and greater dilation of left ventricle dimension with homozygotes [12,34-37]. Fishman, et al. demonstrated that an -174C construct showed lower basal and stimulated expression of the reporter gene in comparison to the -174G construct in HeLa cells. Fishman, et al. also showed an increase in serum IL-6 levels in GG homozygotes in healthy subjects [12]. Italian centenarians, male, GG homozygotes showed higher IL-6 levels [34]. Presence of GG homozygotes for IL-6-174 polymorphism was associated with greater left ventricular dimensions negating the assumption of C allele as determinant of functional status of the DCM [10]. Olivier, et al. identified -174 C/G locus affecting in vitro/in vivo IL-6 production during aging. C allele carriers produced smaller amounts of IL-6 in vitro than C- individuals, and IL-6 C+ individuals had lower plasma levels than C- individuals, and this phenomenon was significant only in men [35]. Though a dichotomy of results has been reported with IL-6 C allele, yet several confounding factors such as age, gender and ever smoking are also revealed suggesting that serum IL-6 levels may not only be due to IL-6 –174 G/C polymorphism.
Clinically patients showed low systolic (P=0.0194) and diastolic (P=0.0011) blood pressure as compared to healthy subjects. Cholesterol (P <0.0001), HDL-C (P <0.0001) and LDL-C (P <0.0001) showed highly significant differences between patients and controls, while, triglycerides and vLDL-C did not show significant difference. A similar clinical was reported in several other reports previously [22,38-40].
Our study concludes that there is strong association of IL-6 -174 G/C polymorphism in the patients with DCM in our population. This is the first study in Pakistani population that showed association between IL-6 -174 G/C gene to DCM. Further in-depth studies are required to confirm our findings, Additionally, IL-6 ELISA analysis would give a more conclusive result for the quantification of IL-6 secreted by the various genotypes.
Conflict of Interest
Authors declare no conflict of interest
Refernces
- Towbin JA, Bowles NE. The failing heart. 2002;415(6868):227-233.
- Jefferies JL, Towbin JA. Dilated cardiomyopathy. Lancet (London, England). 2010;375(9716):752-762.
- Codd MB, Sugrue DD, Gersh BJ, Melton LJ, 3rd. Epidemiology of idiopathic dilated and hypertrophic cardiomyopathy. A population-based study in Olmsted County, Minnesota, 1975-1984. 1989;80(3):564-572.
- Jefferies JL, Towbin JA. Dilated cardiomyopathy. Lancet (London, England). 2010;375(9716):752-762.
- Mann DL, Young JB. Basic mechanisms in congestive heart failure. Recognizing the role of proinflammatory cytokines. 1994;105(3):897-904.
- Aukrust P, Ueland T, Lien E, et al. Cytokine network in congestive heart failure secondary to ischemic or idiopathic dilated cardiomyopathy. The American journal of cardiology. 1999;83(3):376-382.
- Humphries SE, Luong LA, Ogg MS, Hawe E, Miller GJ. The interleukin-6 -174 G/C promoter polymorphism is associated with risk of coronary heart disease and systolic blood pressure in healthy men. European heart journal. 2001;22(24):2243-2252
- Basso F, Lowe GD, Rumley A, McMahon AD, Humphries SE. Interleukin-6 -174G>C polymorphism and risk of coronary heart disease in West of Scotland coronary prevention study (WOSCOPS). Arteriosclerosis, thrombosis, and vascular biology. 2002;22(4):599-604.
- Chang HJ, Chung J, Choi BJ, et al. The origin of proinflammatory cytokines in patients with idiopathic dilated cardiomyopathy. Journal of Korean medical science. 2003;18(6):791-796.
- Adamopoulos S, Kolokathis F, Gkouziouta A, et al. Cytokine gene polymorphisms are associated with markers of disease severity and prognosis in patients with idiopathic dilated cardiomyopathy. 2011;54(1):68-73.
- Song H, Peng Y, Zhou B, et al. Associations between Interleukin-31 Gene Polymorphisms and Dilated Cardiomyopathy in a Chinese Population. Disease markers. 2017;2017:4191365.
- Fishman D, Faulds G, Jeffery R, et al. The effect of novel polymorphisms in the interleukin-6 (IL-6) gene on IL-6 transcription and plasma IL-6 levels, and an association with systemic-onset juvenile chronic arthritis. The Journal of clinical investigation. 1998;102(7):1369-1376.
- Capurso C, Solfrizzi V, D'Introno A, et al. Interleukin 6-174 G/C promoter gene polymorphism and sporadic Alzheimer's disease: geographic allele and genotype variations in Europe. Experimental gerontology. 2004;39(10):1567-1573.
- Li F, Xu J, Zheng J, et al. Association between interleukin-6 gene polymorphisms and rheumatoid arthritis in Chinese Han population: a case-control study and a meta-analysis. Sci Rep. 2014;4:5714-5714.
- Richardson P, McKenna W, Bristow M, et al. Report of the 1995 World Health Organization/International Society and Federation of Cardiology Task Force on the Definition and Classification of cardiomyopathies. 1996;93(5):841-842.
- Mestroni L, Maisch B, McKenna WJ, et al. Guidelines for the study of familial dilated cardiomyopathies. Collaborative Research Group of the European Human and Capital Mobility Project on Familial Dilated Cardiomyopathy. European heart journal. 1999;20(2):93-102.
- Elliott P. Cardiomyopathy. Diagnosis and management of dilated cardiomyopathy. 2000;84(1):106-112.
- Elliott P, Andersson B, Arbustini E, et al. Classification of the cardiomyopathies: a position statement from the European Society Of Cardiology Working Group on Myocardial and Pericardial Diseases. European heart journal. 2008;29(2):270-276.
- Carlson RV, Boyd KM, Webb DJ. The revision of the Declaration of Helsinki: past, present and future. Br J Clin Pharmacol. 2004;57(6):695-713.
- Scheller J, Chalaris A, Schmidt-Arras D, Rose-John S. The pro- and anti-inflammatory properties of the cytokine interleukin-6. Biochimica et biophysica acta. 2011;1813(5):878-888.
- Yamauchi-Takihara K, Kishimoto T. Cytokines and their receptors in cardiovascular diseases--role of gp130 signalling pathway in cardiac myocyte growth and maintenance. Int J Exp Pathol. 2000;81(1):1-16.
- Liaquat A, Asifa GZ, Zeenat A, Javed Q. Polymorphisms of tumor necrosis factor-alpha and interleukin-6 gene and C-reactive protein profiles in patients with idiopathic dilated cardiomyopathy. Annals of Saudi medicine. 2014;34(5):407-414.
- Liaquat A, Shauket U, Ahmad W, Javed Q. The tumor necrosis factor-alpha -238G/A and IL-6 -572G/C gene polymorphisms and the risk of idiopathic dilated cardiomyopathy: a meta-analysis of 25 studies including 9493 cases and 13,971 controls. Clinical chemistry and laboratory medicine. 2015;53(2):307-318.
- Olomolaiye O, Wood NA, Bidwell JL. A novel NlaIII polymorphism in the human IL-6 promoter. European journal of immunogenetics : official journal of the British Society for Histocompatibility and Immunogenetics. 1998;25(2-3):267.
- Capurso C, Solfrizzi V, D'Introno A, et al. Interleukin 6-174 G/C promoter gene polymorphism and sporadic Alzheimer's disease: geographic allele and genotype variations in Europe. Experimental gerontology. 2004;39(10):1567-1573.
- Jenny NS, Tracy RP, Ogg MS, et al. In the elderly, interleukin-6 plasma levels and the -174G>C polymorphism are associated with the development of cardiovascular disease. Arteriosclerosis, thrombosis, and vascular biology. 2002;22(12):2066-2071.
- Bruunsgaard H, Christiansen L, Pedersen AN, Schroll M, Jorgensen T, Pedersen BK. The IL-6 -174G>C polymorphism is associated with cardiovascular diseases and mortality in 80-year-old humans. Experimental gerontology. 2004;39(2):255-261.
- Panoulas VF, Stavropoulos-Kalinoglou A, Metsios GS, et al. Association of interleukin-6 (IL-6)-174G/C gene polymorphism with cardiovascular disease in patients with rheumatoid arthritis: the role of obesity and smoking. 2009;204(1):178-183.
- Ravaglia G, Forti P, Maioli F, et al. Associations of the -174 G/C interleukin-6 gene promoter polymorphism with serum interleukin 6 and mortality in the elderly. 2005;6(6):415-423.
- Brull DJ, Montgomery HE, Sanders J, et al. Interleukin-6 gene -174g>c and -572g>c promoter polymorphisms are strong predictors of plasma interleukin-6 levels after coronary artery bypass surgery. Arteriosclerosis, thrombosis, and vascular biology. 2001;21(9):1458-1463.
- Kilpinen S, Hulkkonen J, Wang XY, Hurme M. The promoter polymorphism of the interleukin-6 gene regulates interleukin-6 production in neonates but not in adults. European cytokine network. 2001;12(1):62-68
- Rea IM, Ross OA, Armstrong M, et al. Interleukin-6-gene C/G 174 polymorphism in nonagenarian and octogenarian subjects in the BELFAST study. Reciprocal effects on IL-6, soluble IL-6 receptor and for IL-10 in serum and monocyte supernatants. Mechanisms of ageing and development. 2003;124(4):555-561.
- Jones KG, Brull DJ, Brown LC, et al. Interleukin-6 (IL-6) and the prognosis of abdominal aortic aneurysms. 2001;103(18):2260-2265.
- Bonafe M, Olivieri F, Cavallone L, et al. A gender--dependent genetic predisposition to produce high levels of IL-6 is detrimental for longevity. European journal of immunology. 2001;31(8):2357-2361.
- Olivieri F, Bonafe M, Cavallone L, et al. The -174 C/G locus affects in vitro/in vivo IL-6 production during aging. Experimental gerontology. 2002;37(2-3):309-314.
- Plenz G, Eschert H, Erren M, et al. The interleukin-6/interleukin-6-receptor system is activated in donor hearts. Journal of the American College of Cardiology. 2002;39(9):1508-1512.
- Plenz G, Song ZF, Tjan TD, et al. Activation of the cardiac interleukin-6 system in advanced heart failure. European journal of heart failure. 2001;3(4):415-421.
- Banerjee I, Pandey U, Hasan OM, Parihar R, Tripathi V, Ganesh S. Association between inflammatory gene polymorphisms and coronary artery disease in an Indian population. Journal of thrombosis and thrombolysis. 2009;27(1):88-94.
- Nakayama T, Asai S, Sato N, Soma M. Genotype and haplotype association study of the STRK1 region on 5q12 among Japanese: a case-control study. 2006;37(1):69-76.
- Saleheen D, Bukhari S, Haider SR, et al. Association of phosphodiesterase 4D gene with ischemic stroke in a Pakistani population. 2005;36(10):2275-2277.


 Impact Factor: * 3.5
Impact Factor: * 3.5 CiteScore: 2.9
CiteScore: 2.9  Acceptance Rate: 14.80%
Acceptance Rate: 14.80%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks