JAL-TA9 Inhibits Aggregation of hPrP180-192 through the Cleavage Reaction
Article Information
Rina Nakamura1,2, Motomi Konishi3, Yuko Sakaguchi4, Yusuke Hatakawa5, Akiko Tanaka5, Toshiyasu Sakane5, Motoaki Saito2, Toshifumi Akizawa1,2,*
1O-Force Co. Ltd., 3454 Irino, Kuroshio-cho, Hata-gun, Kochi 789-1931, Japan
2Laboratory of Pharmacology, School of Medicine, Kochi University, Kohasu, Oko-cho, Nankoku, Kochi, 783-8505, Japan
3Laboratory of Clinical Analytical Chemistry, Faculty of Pharmaceutical Sciences, Setsunan University, 45-1 Nagaotoge-cho, Hirakata, Osaka 573-0101, Japan
4Department of Pharmacy, Japanese Red Cross Kyoto Daiichi Hospital, 15-749 Honmachi, Higashiyama-ku, Kyoto, Kyoto, 605-0981, Japan
5Pharmaceutical Technology, Kobe Pharmaceutical University, 4-19-1, Motoyamakita, Higashinada, Kobe, 658-8558, Japan
*Corresponding author: Toshifumi Akizawa, Laboratory of Clinical Analytical Chemistry, Faculty of Pharmaceutical Sciences, Setsunan University, 45-1 Nagaotoge-cho, Hirakata, Osaka, Japan
Received: 11 May 2020; Accepted: 18 May 2020; Published: 22 May 2020
Citation:
Rina Nakamura, Motomi Konishi, Yuko Sakaguchi, Yusuke Hatakawa, Akiko Tanaka, Toshiyasu Sakane, Motoaki Saito, and Toshifumi Akizawa. JAL-TA9 inhibits aggregation of hPrP180-192 through the cleavage reaction. Journal of Pharmacy and Pharmacology Research 4 (2020): 23-33.
View / Download Pdf Share at FacebookAbstract
We identified that JAL-TA9, a 9-mer synthetic peptide derived from Tob1 protein, possesses catalytic activity and cleaves Aβ42. We also determined that the C-terminal end of Prion protein (PrP) is an essential region for aggregation of PrP using PrP fragment peptides. We investigated the effect of JAL-TA9 on hPrP180-192, which is a fragment peptide derived from human prion protein. Conformational studies were performed using CD analysis. The cleavage reaction was identified using HPLC and MS analyses. JAL TA9 inhibited the formation of aggregated hPrP180-192 and decreased the aggregated hPrP180-192 level through the cleavage reaction. These results indicate that JAL-TA9 could be a promising candidate for developing novel drugs against prion diseases and Alzheimer’s disease.
Keywords
JAL-TA9; catalytide; synthetic hydrolase peptide; synthetic peptide; prion protein; prion fragment peptide; aggregation; CD spectra; MS analysis
JAL-TA9 articles, catalytide articles, synthetic hydrolase peptide articles, synthetic peptide articles, prion protein articles, prion fragment peptide articles, aggregation articles, CD spectra articles, MS analysis articles
Article Details
Introduction
Aggregation is thought to be the critical event in protein misfolding disorders (PMDs), such as prion diseases, Alzheimer’s disease (AD), Parkinson’s disease (PD), and amyotrophic lateral sclerosis (ALS) for inducing toxicity against the central nervous system or peripheral organs. Therefore, many natural compounds and chemical molecules have been tested for developing an effective inhibitor that could inhibit the aggregation of these proteins. To this date, no such efficient therapy that could inhibit the progression of PMDs has been reported [1]. Prion protein (PrP) is a cell-surface glycoprotein that has been associated with the pathogenesis of a range of neurodegenerative disorders which are caused due to the conversion of the cellular prion protein (PrPC) to its protease-resistant misfolded isoform, scrapie prion protein (PrPSc) [2-6]. The underlying mechanisms of conversion and aggregation of PrP remain unexplored, owing to their solubility and aggregation in vitro. In recent years, many researchers have used the fragment peptides of human PrP (hPrP) to understand its physical and physiological properties [7-13]. Structural studies of fragment peptides using nuclear magnetic resonance (NMR) and circular dichroism (CD) have shown that the C-terminal domain plays an important role in the pathogenic properties of PrPSc [14-21]. Furthermore, the C-terminal region, which is composed of two α-helices, two glycosylation sites, and a glycosylphosphatidylinositol anchor inside a folded region [22-23] shows high resistance to proteases such as MT1-MMP, MT3-MMP, and MMP-7. As only MMP can cleave hPrP180-192 with one cleavage site [8], we hypothesized that hPrP180-192 could be an important sequence for the aggregation of PrP and might prove useful in identifying a potent inhibitor of PrP aggregation.
We recently showed that a synthetic 9-mer peptide, JAL-TA9 (YKGSGFRMI), possesses proteolytic activity and cleaves amyloid-β protein (Aβ42) at multiple cleavage sites, especially on the central region. We also proposed that atermed Catalytide (Catalytic peptide) as the generally name of peptide possessing proteolytic activity [24-27]. JAL-TA9 is a fragment peptide from the Box A region of Tob1 protein, which is a highly conserved homology domain of Tob/BTG family of proteins which comprises of Tob1, Tob2, BTG1, BTG2, BTG3/ANA, and BTG4. These proteins possess antiproliferative activities and are involved in the regulation of tumorigenesis. However, there have been no reports about the role of Box A region of Tob/BTG family [28-33]. In this study, we evaluated the effect of JAL-TA9 on the conformational changes and aggregation of hPrP180-192 using CD and HPLC.
Materials and Methods
Materials
Trifluoroacetic acid (TFA, peptide synthesis grade), diethyl ether, acetonitrile (CH3CN, HPLC grade), 2.0 MN, N-diisopropylethylamine/N-methylpyrrolidone (DIEA), and 2-amino-2-hydroxymethyl-1,3-propanediol (Tris) were purchased from Wako Pure Chemical Industries Ltd., Inc. (Japan). Thioanisole was obtained from Nacalai Tesque (Japan), and 1,2-ethanedithiol was obtained from Kanto Chemical Co., Inc. (Japan). HCl was purchased from Kishida Chemical Co., Ltd. (Japan). Piperidine was obtained from Sigma-Aldrich Japan. O-(7-Azabenzo-triazol-1-yl)-N, N’, N', N-tetra-methyluronium hexafluorophosphate (HATU), N’, N-dimethylformamide, and dichloromethane were purchased from Watanabe Chemical Ind., Ltd. (Japan). Fluorenylmethoxycarbonyl (Fmoc)-L-amino acid preloaded resin and Fmoc-L-amino acid were purchased from AAPPTec (USA). Milli-Q water was used for all experiments.
Methods
Chemical synthesis of the peptides: Peptides were synthesized from Fmoc-protected L-amino acid derivatives according to the method described by Kojima et al [8]. Fragment peptide hPrP180-192: VNITIKQHTVTTT corresponding to amino acid sequences from human prion protein and fragment peptide JAL-TA9: YKGSGFRMI corresponding to amino acid sequences from the Box A region of Tob1 protein were synthesized using an automated Applied Biosystems (ABI, USA) peptide synthesizer, model 433A (0.1 mmol scale with preloaded resin). After performing deprotection according to the manufacturer’s protocol, the peptides were purified using reversed-phase HPLC (Capcell Pak C18 column, SG, 10 mm i.d. × 250 mm; Shiseido Co., Ltd., Japan) at the flow rate of 3.0 mL/min with a linear elution gradient ranging from 0.1 % TFA to 50 % CH3CN containing 0.1 % TFA over 30 min. The primary peak was collected and lyophilized. The purity of the synthetic peptides and the progress of the enzymatic reaction were confirmed by analytical reversed-phase HPLC (Capcell Pak C18 column, MGII, 4.6 mm i.d. × 150 mm; Shiseido Co. Ltd., Japan) at a flow rate of 1.0 mL/min with a linear elution gradient from 0.1 % TFA to 70 % CH3CN containing 0.1 % TFA. The column eluate was monitored with a photodiode-array detector (SPD-M20A; Shimadzu, Japan). Each purified peptide was characterized by ESI-MS using a Qstar Elite Hybrid LC/MS/MS system (ABI, USA).
CD analysis: hPrP180-192 and JAL-TA9 were dissolved in 10 mM Tris-HCl to obtain a final concentration of 0.025 mM. The pH of Tris-HCl was adjusted to 7.5 using HCl. Reaction mixture of hPrP180-192 and JAL-TA9 were incubated at 37 °C. CD analysis was performed using a J-805 spectropolarimeter (JASCO, Japan). The CD spectra were measured using a 0.2 cm path-length quartz cell. Signals were collected from 190 nm to 260 nm at approximately 26 °C with a scan rate of 10 nm/min, and a response time of 4 s. Each spectrum comprises of four accumulated scans.
Analysis of proteolytic activity and determination of cleavage sites: 0.025 mM hPrP180-192 was incubated with and without the 0.2 mM JAL-TA9 in 10 mM Tris-HCl (pH 7.5) at 37 °C. Twenty microliters of each reaction mixture was analyzed in a time-dependent manner using the analytical HPLC system described in section 2.2.2. For the peak collection, 40 μL of reaction mixture was loaded into the same HPLC system, and the peak fractions monitored at 220 nm were collected into microtubes (Eppendorf Safe-Lock Tubes, 1.5 mL). After lyophilization, the appropriate quantity of 36 % CH3CN containing 0.1 % HCOOH was determined based on the chromatographic peak height and the two reagents were mixed with the help of an automatic mixer. The cleavage sites were determined by ESI-MS using the flow injection method with 70 % CH3CN containing 0.1 % HCOOH on a Qstar Hybrid LC-MS/MS system (ABI). The flow rate was set at 0.1 mL/min.
Results and Discussion
Effect of JAL-TA9 on conformational change (CD) in hPrP180-192
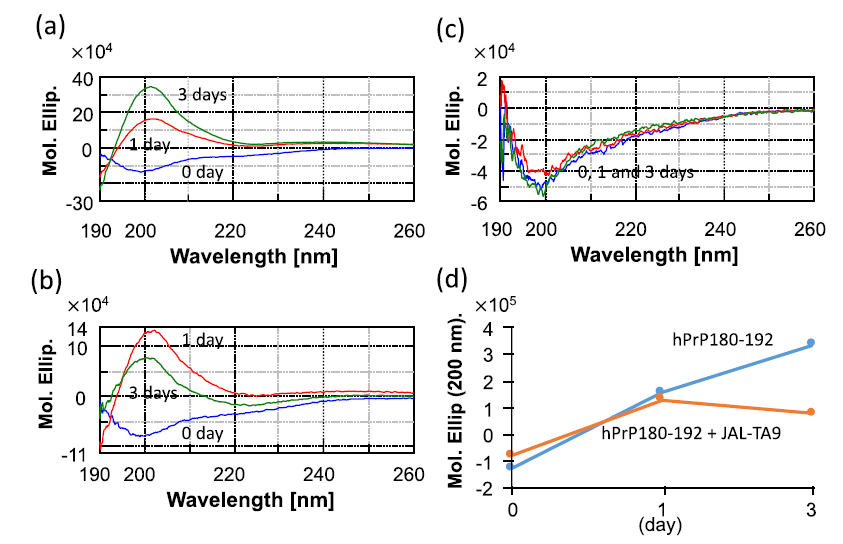
We first examined whether JAL-TA9 affects the conformational change in hPrP180-192 (from a random coil structure to a β-sheet structure) and its oligomerization under physiological conditions using CD spectra. The aggregated hPrP180-192 was sedimented in a buffer [34-35]. The reaction mixture was thoroughly mixed using a vortex mixer and was then suspended to measure the CD spectra. The CD spectra of 0.025 mM hPrP180-192 in the absence and presence of 0.025 mM JAL-TA9 were measured at wavelengths ranging from 190 nm to 260 nm in a 0.2 cm cell on the following days: 0, 1, and 3 (Figure 1). The CD spectra of hPrP180-192 measured on 0 day showed a negative band at 195 nm, which corresponds to a typical random coil structure [36-37]. After one day, hPrP180-192 switched from a random coil structure to a β-sheet structure and gave a positive band at approximately 200 nm. Furthermore, after three days, the positive band was increased (Figure 1a). The CD spectra of hPrP180-192 presence of JAL-TA9 also showed a random coil structure on day 0 and switched to β-sheet on 1st day (Figure 1b). However, the quantity of hPrP180-192 folded in the β-sheet conformation decreased on the 3rd day when compared with 1st day. The spectra of JAL-TA9 did not show any change on day 0, 1, and 3 (Figures 1b and c).
Figure 1: Time dependency of the structural changes of hPrP180-192: (a) CD spectra for 0.025 mM of hPrP180-192 incubated for 0, 1, and 3 days; (b) CD spectra for 0.025 mM of hPrP180-192 co-incubated with 0.025 mM of JAL-TA9 for 0, 1, and 3 days; (c) CD spectra for 0.025 mM of JAL-TA9 for 0, 1, and 3 days; (d) Molar ellipticity at 200 nm plotted for CD spectra of hPrP180-192 with or without JAL-TA9.
We compared the molar ellipticity of the samples at the wavelength of 200 nm, which corresponds to the β-sheet structure [34-35]. No differences were observed on day 0 and 1. On the 3rd day, we observed differences between the molar ellipticity of hPrP180-192 in the presence and absence of JAL-TA9 (Figure 1d). These results indicate that the quantity of hPrP180-192 in the β-sheet conformation increased in a time-dependent manner and was suppressed in the presence of JAL-TA9 (Figures 1c and d). Furthermore, JAL-TA9 was able to affect hPrP180-192 post the conformational changes (from random coil to β-sheet structure).
Proteolytic activity of JAL-TA9 against hPrP180-192
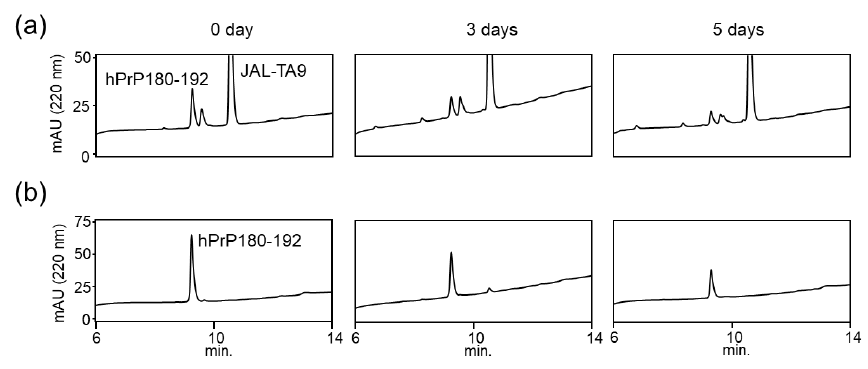
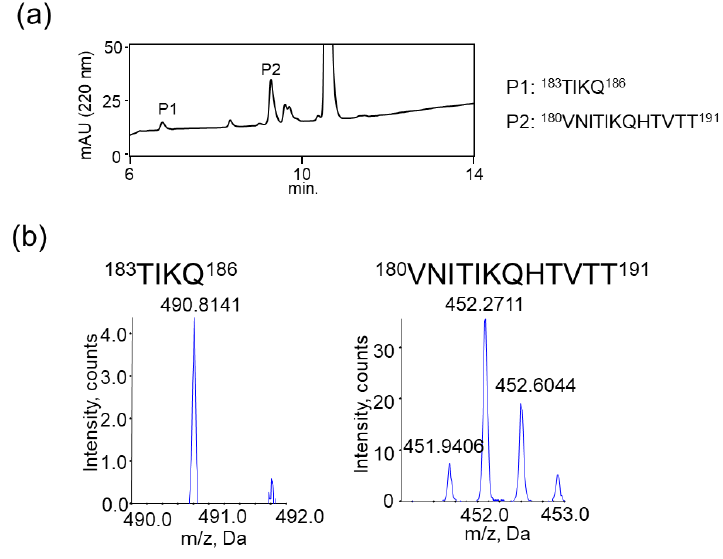
We also examined the proteolytic activity of JAL-TA9 against hPrP180-192 to understand how JAL-TA9 suppresses the β-sheet structure of hPrP180-192. The proteolytic activity was analyzed as per the protocol described in our previous studies [24-27]. The reaction mixture containing 0.025 mM of hPrP180-192 and 0.2 mM of JAL-TA9 was co-incubated in 10 mM Tris-HCl buffer (pH 7.5) at 37 °C, and 20 µL of that reaction mixture was analyzed by analytical (HPLC) from day 0 day to day 5 (Figures S1 and 2). To identify the cleavage site, 40 µL of the reaction mixture was analyzed and corrected all peaks appeared on 5 days and then applied by MS analysis (Figure S1). The chromatograms revealed a slight decrease in hPrP180-192 levels due to its aggregation; however, no new peaks were observed (Figures 2a and S1). In contrast, new peaks appeared, and the peak height of hPrP180-192 showed a decrease in the presence of JAL-TA9 (Figures 2 b and S2). All newly appearing peaks were collected and analyzed using MS analysis. P1 and P2 were identified as hPrP183-186 and hPrP180-191, respectively (Figure 3). The other peaks were identified as fragment peptides of JAL-TA9 produced by auto-degradation (Figure S2). These data indicated that JAL-TA9 cleaved hPrP180-192 and led to the suppression of conformational changes in hPrP180-192.
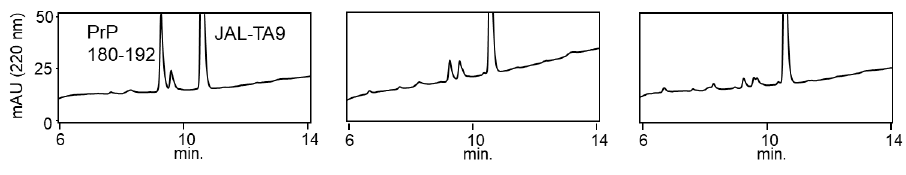
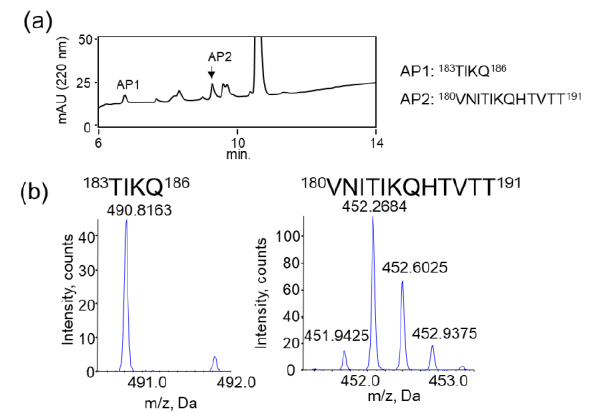
We examined the ability of JAL-TA9 to cleave the aggregated hPrP180-192. We have previously reported that hPrP180-192 completely changes to β-sheet conformation on day 3, and the fluorescence intensity (using the Thioflavin-T assay) increases in a time-dependent manner until day 4 [35]. Based on these observations, we incubated 0.05 mM hPrP180-192 in 10 mM Tris-HCl buffer (pH 7.5) at 37 °C for 3 days to obtain aggregated hPrP180-192 and used it as a substrate. Twenty microliters of the reaction mixture made of JAL-TA9 and aggregated hPrP180-192 was analyzed every day for five days. The chromatograms changed continuously during the five days, and the peak height of hPrP180-192 decreased in a time-dependent manner (Figure 4). By MS analysis, AP1 and AP2 were identified as hPrP 183-186 and hPrP180-191, respectively (Figures 5 and S3). These results are similar to those observed for non-aggregated hPrP180-192 (Figure 3). The cleavage sites on non-aggregated hPrP180-192 and aggregated hPrP180-192 are summarized in (Figure 6).
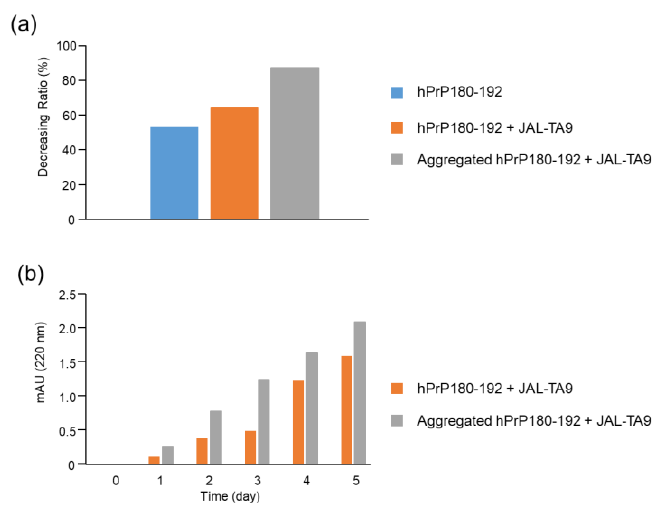
Figure 7: Decreasing ratios of hPrP180-192 and increasing position of the TIKQ. (a) The decreasing ratio was calculated by estimating the peak height of the chromatogram on day 0 and day 5 using HPLC. The chromatogram of hPrP180-192, hPrP180-192 + JAL-TA9, and aggregated hPrP180-192 + JAL-TA9 are shown in Figure 2b (supplementary 1b), 2a (supplementary 1a), and Figure 4 (supplementary 1c). Formula: Peak height on day 5 / Peak height on day 0 × 100 (b) Increasing position of the TIKQ fragment peptide from hPrP180-192 cleaved by JAL-TA9.
The decreasing ratio of hPrP180-192, hPrP180-192 co-incubated with JAL-TA9, and aggregated hPrP180-192 co-incubated with JAL-TA9 is shown in Figure 7a. The decreasing ratio of aggregated hPrP 180-192 co-incubated with JAL-TA9 was higher than that of non-aggregated hPrP180-192. Furthermore, we compared the peak heights of P1 and AP1, both of which correspond to hPrP 183-186. P1 produced from non-aggregated hPrP180-192 showed a small increase from day 1 to day 3; but showed a large increased after four days. On the other hand, for aggregated hPrP180-192, the increment of AP1 was unchanged from day 1 to day 5 (Figure 7). Taken together, these results indicate that JAL-TA9 interacts with hPrP180-192 after aggregation, and then proceeds to cleave it. These data suggest that JAL-TA9 has tendency to cleave the aggregated hPrP180-192 as well as Ab42 [24, 26].
Conclusion
JAL-TA9, the shorter hydrolase peptide can cleave aggregated hPrP180-192 and inhibits the formation of aggregated hPrP180-192 via cleavage reaction. These results suggest that JAL-TA9 could be a promising candidate for designing new strategic peptide drugs against prion diseases.
Patents
- Yamamoto and T. Akizawa, Novel peptide, Patent Pending, (2016) No. US62/275,599.
- Yamamoto and T. Akizawa, Novel hydrolase-like peptide and its use, (2016) No. JP2016-068496
Supplementary Materials
The following are available online at www.mdpi.com/xxx/s1, Figure S1: Time dependent analysis of hPrP180-192 from 0 day to 5 days, Figure S2: Cleavage reaction of hPrP180-192 by JAL-TA9 Figure S3: Cleavage reaction of aggregated hPrP180-192 by JAL-TA9.
Author Contributions
Conceptualization, T.A. and R.N.; methodology, R.N., Y.S. and Y.H.; validation, T.A. and M.K.; formal analysis, R.N., M.K. and T.A.; investigation, R.N. and M.K.; data curation, R.N., M.T and Y.S.; writing—original draft preparation, R.N. and T.A.; writing—review and editing, T.A., M.K., T.S. and M.S; supervision, T.A.; project administration, T.A. and M.S; funding acquisition, T.A. and M.K.
Funding
This work was supported by the Japan Society for the Promotion of Science (JSPS) Grants-in-Aid for Scientific Research (KAKENHI) Program, Grant Number 15K07908.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Dhouafli Z, Cuanalo-Contreras K, Hayouni EA, et al. Inhibition of protein misfolding and aggregation by natural phenolic compounds. Mol. Life Sci 75 (2018): 3521-3538.
- Prusiner SB. Novel proteinaceous infectious particles cause scrapie. Science 216 (1982): 136-144.
- Prusiner SB. Molecular biology of prion diseases. Science 252 (1991): 1515-1522.
- Bendheim PE, Brown HR, Rudelli RD, et al. Nearly ubiquitous tissue distribution of the scrapie agent precursor protein. Neurology (2012).
- Takada LT, Geschwind MD. Prion diseases. Semin Neurol 95 (2013): 348-356.
- Schmitz M, Cramm M, Llorens F, et al. Application of an in vitro-amplification assay as a novel pre-screening test for compounds inhibiting the aggregation of prion protein scrapie. Rep 6 (2016): 1-8.
- Kojima A, Konishi M, Akizawa T. Metal-Binding Ability of His-Containing Fragment Peptides Originating from Prion Protein Studied by Column Switch HPLC. Bunseki Kagaku 59 (2010): 701-707.
- Kojima A, Konishi M, Akizawa T. Prion fragment peptides are digested with membrane type matrix metalloproteinases and acquire enzyme resistance through Cu2+-binding. Biomolecules 4 (2014).
- Kojima A, Sakaguchi Y, Toyoda H, et al. C-terminal Region of the hPrP Can Be the Core for the Structural Conversion and Aggregation. Peptide Science 52 (2015): 122-125.
- Bocharova OV, Breydo L, Salnikov VV, et al. Copper(II) inhibits in vitro conversion of prion protein into amyloid fibrils. Biochemistry 44 (2005): 6776-6787.
- Jones CE, Abdelraheim SR, Brown DE, et al. Preferential Cu2+ coordination by His96 and His 111 induces β-sheet formation in the unstructured amyloiodogenic region of the prion protein. Biol. Chem 279 (2004): 32018-32027.
- Valensin D, Padula EM, Hecel A, et al. Specific binding modes of Cu(I) and Ag(I) with neurotoxic domain of the human prion protein. Inorg. Biochem 155 (2016): 26-35.
- Jobling MF, Stewart LR, White AR, et al. Prion Disease: What Is the Neurotoxic Molecule. Neurochem 73 (1999): 1557-1565.
- Barbara T, Pasquale P, AntoniaDe C, et al. The human prion protein α2 Helix: A thermodynamic study of its conformational preferences. Proteins: Structure, Function, and Bioinformatics 59 (2005): 72-79.
- Colacino S, Tiana G, Broglia RA, et al. The determinants of stability in the human prion protein: Insights into folding and misfolding from the analysis of the change in the stabilization energy distribution in different conditions. Proteins Struct. Funct. Genet 62 (2006): 698-707.
- Dima RI, Thirumalai D. Exploring the propensities of helices in PrPC to form β sheet using NMR structures and sequence alignments. J ( 2002).
- Dima RI, Thirumalai D. Probing the instabilities in the dynamics of helical fragments from mouse PrPC. Natl. Acad. Sci (2004).
- Lu X, Wintrode PL, Surewicz WK. Β-Sheet Core of Human Prion Protein Amyloid Fibrils As Determined By Hydrogen/Deuterium Exchange. Natl. Acad. Sci. U. S. A 104 (2007): 1510-1515.
- Singh J, Udgaonkar JB. Molecular Mechanism of the Misfolding and Oligomerization of the Prion Protein: Current Understanding and Its Implications. Biochemistry (2015).
- Ronga L, Palladino P, Saviano G, et al. Structural characterization of a neurotoxic threonine-rich peptide corresponding to the human prion protein α2-helical 180-195 segment and comparison with full-length α2-helix-derived peptides. Pept. Sci (2008).
- Thompson A, White AR, McLean C, et al. Amyloidogenicity and neurotoxicity of peptides corresponding to the helical regions of PrP(c). Neurosci. Res 62 (2000): 293-301.
- Harris DA, Huber MT, van Dijken P, et al. Processing of a Cellular Prion Protein: Identification of N- and C-Terminal Cleavage Sites. Biochemistry 32 (1993): 1009-1016.
- Zahn R, Liu A, Lührs T, et al. NMR solution structure of the human prion protein. Natl. Acad. Sci. U. S. A 97 (2000): 145-150.
- Nakamura R, Konishi M, Taniguchi M, et al. The discovery of shorter synthetic proteolytic peptides derived from Tob1 protein. Peptides (2019): 0-1.
- Nakamura R, Konishi M, Higashi Y, et al. Comparison of the catalytic activities of 5-mer synthetic peptides derived from Box A region of Tob/BTG family proteins against the amyloid-beta fragment peptides. Mol. Med 6 (2019): 1-4.
- Nakamura R, Konishi M, Hatakawa Y, et al. The Novel Catalytic Peptide, A Synthetic Nona-Peptide (JAL-TA9) Derived from Tob1 Protein, Digests the Amyloid-β Peptide. Royal, Sci 1 (2019): 30-35.
- Hatakawa Y, Nakamura R, Konishi M, et al. Catalytides derived from the Box A region in the ANA/BTG3 protein cleave amyloid-β fragment peptide. Heliyon 5 (2019):
- Winkler GS. The mammalian anti-proliferative BTG/Tob protein family. Cell. Physiol 222 (2010): 66-72.
- Schulze-Topphof U, Casazza S, Varrin-Doyer M, et al. Tob1 plays a critical role in the activation of encephalitogenic t cells in cns autoimmunity. Exp. Med 210 (2013): 1301-1309.
- Chen Y, Wang C, Wu J, et al. BTG/Tob family members Tob1 and Tob2 inhibit proliferation of mouse embryonic stem cells via Id3 mRNA degradation. Biophys. Res. Commun 462 (2015): 208-214.
- Yang X, Morita M, Wang H, et al. Crystal structures of human BTG2 and mouse TIS21 involved in suppression of CAF1 deadenylase activity. Nucleic Acids Res 36 (2008): 6872-6881.
- Matsuda S, Kawamura-Tsuzuku J, Ohsugi M, et al. Tob, a novel protein that interacts with p185erbB2, is associated with anti-proliferative activity. Oncogene (1996).
- Horiuchi M, Takeuchi K, Noda N, et al. Structural basis for the antiproliferative activity of the Tob-hCaf1 complex. Biol. Chem 284 (2009): 13244-13255.
- Sakaguchi Y, Nakamura R, Konishi M, et al. The effects of Cu2+ on conformational changes of hPrP180-192 derived from the C-terminal region of prion protein. BRTE 30 (2019): 1-8.
- Sakaguchi Y, Nakamura R, Konishi M, et al. Effects of Cu2+ on conformational change and aggregation of hPrP180-192 with a V180I mutation of the prion protein. BBRC 514 (2019): 798-802.
- Narashima S, Robert WW. Structural composition of βI-and βII- proteins. Protein Science 12 (2003): 384-388.
- Jie W, Jen T Yang, Chuen-Shang CW. β-II conformation of all-β proteins can be distinguished from inordered from by circular dichroism. Biochem 200 (1992): 359-364.



 Impact Factor: * 2.5
Impact Factor: * 2.5 CiteScore: 2.9
CiteScore: 2.9  Acceptance Rate: 11.01%
Acceptance Rate: 11.01%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 