Outcome and Risk Factors of Overall Survival for Acute Myeloid Leukemia Patients Receiving Allogeneic Hematopoietic Stem Cell Transplantation in First Complete Remission
Article Information
Haiyan Bao1,2,3, Vijay Kumar Kolluri4, Jia Chen1,2,3, Xiao Ma1,2, Yue Han1,2, Chengcheng Fu1,2, Yang Mei5*, Depei Wu1,2,3*
1Jiangsu Institute of Hematology, the First Affiliated Hospital of Soochow University, Suzhou, Jiangsu Province, China
2Institute of Blood and Marrow Transplantation, Soochow University, Suzhou, Jiangsu Province, China
3Collaborative Innovation Center of Hematology, Soochow University, Suzhou, Jiangsu Province, China
4Medical College, Soochow University, Suzhou, Jiangsu Province, China
5College of Biology, Hunan University, Changsha, Hunan Province, China
*Corresponding Authors: Dr. Yang Mei, College of Biology, Hunan University, Changsha, Hunan Province, China
Dr. Depei Wu, Jiangsu Institute of Hematology, the First Affiliated Hospital of Soochow University, Suzhou, Jiangsu Province, China
Received: 17 August 2020; Accepted: 02 September 2020; Published: 01 October 2020
Citation: Haiyan Bao, Vijay Kumar Kolluri, Jia Chen, Xiao Ma, Yue Han, Chengcheng Fu, Yang Mei, Depei Wu. Immune Response in Patients Diagnosed with Non-Muscle Invasive Bladder Cancer Treated with CIMAvax-EGF Concomitant with Intravesical Bacillus Calmette-Guerin. Journal of Cancer Science and Clinical Therapeutics 4 (2020): 369-381.
View / Download Pdf Share at FacebookAbstract
Objective: Acute myeloid leukemia (AML) is the most common acute leukemia in adults. To investigate how we can improve the efficacy of allogeneic hematopoietic stem cell transplantation (HSCT) for AML patients in first complete remission (CR1), we analyze the risk factors of overall survival (OS), and depict gene mutation profiles in the patients.
Materials and Methods: AML patients undergoing allogeneic HSCT in CR1 from January 2011 to January 2015 were retrospectively reviewed. We compared patients’ disease characteristics and survival outcomes between the survivor and non-survivor groups. The study cohort comprised of 135 AML patients.
Results: The OS for all patients was 63.7%. Gene mutations were commonly detected. The genes with mutation incidence > 10% include NRAS, ASXL1, TET2, IDH1/2, C-KIT, CEBPA biallelic, WT1, NPM1, DNMT3A, and FLT3-ITD. The risk factors found in univariate analysis were age, risk group at diagnosis, more than one course to achieve CR1, acute graft versus host disease (aGVHD) grade III-IV, without chronic graft versus host disease (cGVHD), relapse, and IDH1/2 mutation. Multivariate analysis further identified that aGVHD grade III-IV and relapse are independent risk factors of OS, whereas cGVHD is a favorable factor. IDH1/2 mutation may confer an adverse effect on allogeneic HSCT.
Conclusions: Our study suggests that allogeneic HSCT is a curative way for the majority of AML patients in CR1. New strategies to control severe aGVHD and relapse are emerging. Targeted therapy (e.g., IDH 1/2 inhibitors) and manipulation of immune system may contribute to an improved OS in these patients.
Keywords
Acute myeloid leukemia; Allogeneic stem cell transplantation; Overall survival; IDH2
Acute myeloid leukemia articles, Allogeneic stem cell transplantation articles, Overall survival articles, IDH2 articles
Acute myeloid leukemia articles Acute myeloid leukemia Research articles Acute myeloid leukemia review articles Acute myeloid leukemia PubMed articles Acute myeloid leukemia PubMed Central articles Acute myeloid leukemia 2023 articles Acute myeloid leukemia 2024 articles Acute myeloid leukemia Scopus articles Acute myeloid leukemia impact factor journals Acute myeloid leukemia Scopus journals Acute myeloid leukemia PubMed journals Acute myeloid leukemia medical journals Acute myeloid leukemia free journals Acute myeloid leukemia best journals Acute myeloid leukemia top journals Acute myeloid leukemia free medical journals Acute myeloid leukemia famous journals Acute myeloid leukemia Google Scholar indexed journals Allogeneic stem cell transplantation articles Allogeneic stem cell transplantation Research articles Allogeneic stem cell transplantation review articles Allogeneic stem cell transplantation PubMed articles Allogeneic stem cell transplantation PubMed Central articles Allogeneic stem cell transplantation 2023 articles Allogeneic stem cell transplantation 2024 articles Allogeneic stem cell transplantation Scopus articles Allogeneic stem cell transplantation impact factor journals Allogeneic stem cell transplantation Scopus journals Allogeneic stem cell transplantation PubMed journals Allogeneic stem cell transplantation medical journals Allogeneic stem cell transplantation free journals Allogeneic stem cell transplantation best journals Allogeneic stem cell transplantation top journals Allogeneic stem cell transplantation free medical journals Allogeneic stem cell transplantation famous journals Allogeneic stem cell transplantation Google Scholar indexed journals Overall survival articles Overall survival Research articles Overall survival review articles Overall survival PubMed articles Overall survival PubMed Central articles Overall survival 2023 articles Overall survival 2024 articles Overall survival Scopus articles Overall survival impact factor journals Overall survival Scopus journals Overall survival PubMed journals Overall survival medical journals Overall survival free journals Overall survival best journals Overall survival top journals Overall survival free medical journals Overall survival famous journals Overall survival Google Scholar indexed journals IDH2 articles IDH2 Research articles IDH2 review articles IDH2 PubMed articles IDH2 PubMed Central articles IDH2 2023 articles IDH2 2024 articles IDH2 Scopus articles IDH2 impact factor journals IDH2 Scopus journals IDH2 PubMed journals IDH2 medical journals IDH2 free journals IDH2 best journals IDH2 top journals IDH2 free medical journals IDH2 famous journals IDH2 Google Scholar indexed journals Targeted therapy articles Targeted therapy Research articles Targeted therapy review articles Targeted therapy PubMed articles Targeted therapy PubMed Central articles Targeted therapy 2023 articles Targeted therapy 2024 articles Targeted therapy Scopus articles Targeted therapy impact factor journals Targeted therapy Scopus journals Targeted therapy PubMed journals Targeted therapy medical journals Targeted therapy free journals Targeted therapy best journals Targeted therapy top journals Targeted therapy free medical journals Targeted therapy famous journals Targeted therapy Google Scholar indexed journals malignant hematologic articles malignant hematologic Research articles malignant hematologic review articles malignant hematologic PubMed articles malignant hematologic PubMed Central articles malignant hematologic 2023 articles malignant hematologic 2024 articles malignant hematologic Scopus articles malignant hematologic impact factor journals malignant hematologic Scopus journals malignant hematologic PubMed journals malignant hematologic medical journals malignant hematologic free journals malignant hematologic best journals malignant hematologic top journals malignant hematologic free medical journals malignant hematologic famous journals malignant hematologic Google Scholar indexed journals alloreactivity articles alloreactivity Research articles alloreactivity review articles alloreactivity PubMed articles alloreactivity PubMed Central articles alloreactivity 2023 articles alloreactivity 2024 articles alloreactivity Scopus articles alloreactivity impact factor journals alloreactivity Scopus journals alloreactivity PubMed journals alloreactivity medical journals alloreactivity free journals alloreactivity best journals alloreactivity top journals alloreactivity free medical journals alloreactivity famous journals alloreactivity Google Scholar indexed journals haploidentical donors articles haploidentical donors Research articles haploidentical donors review articles haploidentical donors PubMed articles haploidentical donors PubMed Central articles haploidentical donors 2023 articles haploidentical donors 2024 articles haploidentical donors Scopus articles haploidentical donors impact factor journals haploidentical donors Scopus journals haploidentical donors PubMed journals haploidentical donors medical journals haploidentical donors free journals haploidentical donors best journals haploidentical donors top journals haploidentical donors free medical journals haploidentical donors famous journals haploidentical donors Google Scholar indexed journals molecular pathophysiology articles molecular pathophysiology Research articles molecular pathophysiology review articles molecular pathophysiology PubMed articles molecular pathophysiology PubMed Central articles molecular pathophysiology 2023 articles molecular pathophysiology 2024 articles molecular pathophysiology Scopus articles molecular pathophysiology impact factor journals molecular pathophysiology Scopus journals molecular pathophysiology PubMed journals molecular pathophysiology medical journals molecular pathophysiology free journals molecular pathophysiology best journals molecular pathophysiology top journals molecular pathophysiology free medical journals molecular pathophysiology famous journals molecular pathophysiology Google Scholar indexed journals Marrow Transplantation articles Marrow Transplantation Research articles Marrow Transplantation review articles Marrow Transplantation PubMed articles Marrow Transplantation PubMed Central articles Marrow Transplantation 2023 articles Marrow Transplantation 2024 articles Marrow Transplantation Scopus articles Marrow Transplantation impact factor journals Marrow Transplantation Scopus journals Marrow Transplantation PubMed journals Marrow Transplantation medical journals Marrow Transplantation free journals Marrow Transplantation best journals Marrow Transplantation top journals Marrow Transplantation free medical journals Marrow Transplantation famous journals Marrow Transplantation Google Scholar indexed journals
Article Details
1. Introduction
Acute myeloid leukemia (AML) is the most common acute leukemia in adults with clinically and biologically heterogeneous characteristics [1]. Allogeneic hematopoietic stem cell transplantation (HSCT) is a promising therapy for both benign and malignant hematologic disorders. With advances in infection control, enhancement of hematopoiesis recovery, and patient education on personal care, transplant-related mortality (TRM) has been greatly lowered [2]. Due to the improved strategy on suppression of alloreactivity, haploidentical donors are also becoming an alternative choice to perform allogeneic HSCT in AML patients [3]. A growing number of AML patients are undergoing allogeneic HSCT. As tremendous advances made in sequencing technologies, especially the next generation sequencing (NGS), the discovery in the molecular pathophysiology of AML comes to a new era [4]. Numerous genes are found to be recurrently mutated in AML. Molecular screening can help us to identify gene mutations affecting prognostic stratification and disease monitoring, and further guide the following treatment.
Here, we report the outcome and risk factors on long-term overall survival (OS) of AML patients undergoing allogeneic SCT in first complete remission (CR1), together with their gene mutation profile in our transplantation center. Their disease characteristics and survival data were retrospectively reviewed and compared between the survivor and non-survivor groups with a median follow-up of up to 7 years.
2. Materials and Methods
2.1 Study population
One hundred and thirty-five patients diagnosed with de novo acute myeloid leukemia (AML) who underwent allogeneic HSCT between January 2011 and January 2015 in the Institute of Blood and Marrow Transplantation, Soochow University, China, were retrospectively reviewed in this study. Patients were scored into two groups according to their survival status: the survivor group and the non-survivor group. Clinical data and molecular pathophysiology were retrieved from the medical records in our unit. This study was approved by the institutional ethics committee of the First Affiliated Hospital of Soochow University.
2.2 Treatment before transplantation
The routine treatment for induction in our department was a standard first-line “7 + 3” regimen composed of daunorubicin (60-90mg/m2 for 3 days) or idarubicin (12mg/m2 for 3 days) and cytarabine (100mg/m2 for 7 days). After achieving CR1, patients received consolidation chemotherapies with a similar “7 + 3” regimen consisting of an alternative anthracycline or anthraquinone agents, or chemotherapy containing intermediate/high-dose cytarabine (Ara-C), at a dose of 1-2g/m2 q12h for 3 days per cycle. The number of cycles patients received prior to transplantation was based on AML risk category, minimal residual disease (MRD), performance status, and financial situation.
2.3 Conditioning regimen and stem cell harvest
The conditioning regimen for allogeneic HSCT was myeloablative. Patients received a conditioning regimen consisting of cytarabine (2 g/m2/day q12h for 1 day), busulfan (0.8 mg/kg/day q6h for 3 days, intravenously), and cyclophosphamide (1.8 g/m2/day for 2 days). Patients who had an HLA-matched sibling donor underwent an HLA-matched sibling transplant. Patients who failed to find an HLA-matched sibling donor received HLA-matched unrelated HSCT or HLA haploidentical HSCT. Donors received granulocyte colony-stimulating factor (G-CSF; 5 μg/kg/day for 5 days, subcutaneously) to mobilize hematopoietic stem cells. Bone marrow (BM)-derived allograft was preferred for haploidentical HSCT. If BM-derived stem cells were not enough, peripheral blood stem cells (PBSCs) would be collected in the coming days. The target CD34+ stem cell dose was 2 × 106/kg per recipient’s body weight. For an HLA-matched sibling or unrelated donor, PBSCs were recommended. BM-derived stem cells were collected through BM aspiration under general anesthesia. Peripheral blood allografts were harvested by cell apheresis.
2.4 Graft versus host disease (GVHD) prophylaxis
Cyclosporine was used to prevent GVHD at a dose of 3 mg/kg/day by continuous infusion, from day -1 to the day when patients could tolerate an oral agent. The range of target serum concentration for cyclosporine was 200-300 ng/ml. Fractional methotrexate (MTX) doses were also given on days +1, +3, +6 at a dose of 15, 10, 10 mg/m2, respectively. If the patient received unrelated or haploidentical donor allografts, an additional dose of MTX was administrated on day +11 (10 mg/m2). Rabbit anti-thymocyte globulin (ATG; 2.5 mg/kg/day, starting on day -5 to day -2), together with orally intake of mycophenolate mofetil (MMF; 30 mg/kg/day from day -9 to day +30) was added to the latter regimen additionally.
2.5 Supportive care
Selective gut decontamination (albendazole and levofloxacin), ganciclovir, fluconazole, and trimethoprim-sulfamethoxazole were administered prior to transplantation to prevent opportunistic infection. Heparin and prostaglandin E1 were used to prevent sinusoidal obstruction syndrome (SOS). G-CSF and thrombopoietin (TPO) was started on day +7 to accelerate the recovery of hematopoiesis. Supportive care of irradiated red blood cells and platelets transfusions were considered to maintain a hemoglobin concentration > 60 g/L and platelet count > 20 × 109/L.
2.6 Molecular examinations
Chimerism of donor cells in recipient’s peripheral blood was calculated by short tandem repeat-polymerase chain reaction (STR-PCR) weekly after transplantation. Cytomegalovirus (CMV) and Epstein-Barr virus (EBV) viral load was detected by real-time polymerase chain reaction (RT-PCR). BM aspiration was done monthly to evaluate disease status within 3 months post-SCT, every 3 months for one year, and then yearly for at least 5 years.
BM samples were aspirated and processed under 24-hour unstimulated culture. Karyotype was identified using conventional R-banding assay. At least 20 metaphases were karyotyped per BM sample. Risk stratification of cytogenetics was based on the National Comprehensive Cancer Network (NCCN) guidelines Version 1, 2020 for AML. Genomic DNA was extracted from BM samples using the PurelinkTM Genomic DNA mini kit (Invitrogen, Carlsbad, CA). A panel of recurrently mutated genes in AML, including signal genes (C-KIT, FLT3-ITD, PTPN11, KRAS, NRAS), myeloid transcription factor mutations (CEBPA biallelic, RUNX1), chromatin-modifying genes (ASXL1, EZH2), DNA methylation-associated genes (DNMT3A, TET2, IDH1, IDH2), tumor-suppressor genes (TP53, WT1, PHF6), spliceosome-complex genes (SF3B1, U2AF1), cohesin-complex gene (RAD21) and NPM1, were screened by NGS technique (GENESEE, Nanjing, China).
2.7 Definitions
Neutrophil engraftment was defined as the peripheral blood neutrophil count ≥ 0.5×109/L for 3 consecutive days. Platelet engraftment was defined as the platelet count ≥ 20×109/L for a week without transfusion. The severity of acute GVHD (aGVHD) was graded by Glucksberg-Seattle system [5]. Chronic GVHD (cGVHD) was recorded based on the National Institutes of Health Consensus Development Project [6]. Relapse was defined as blast percentage in BM ≥ 5%, or recurrence of disease-specific fusion genes. CMV and EBV viremias were assured by detection of viral DNA copies.
2.8 Statistical analyses
All statistical calculations were performed using the SPSS version 19.0 for Windows. A student’s t-test was used for continuous variables to compare the age, white blood cells (WBC) count, BM blasts at diagnosis. The Chi-square test was used to compare categorical variables. Overall survival (OS) was defined as the time from disease diagnosis to death or last follow-up. Cox’s hazard model was used to determine risk factors related to mortality. Variables in the univariate analysis (P < 0.1) were included in the regression model. Independent risk factors for mortality were obtained in the multivariate analysis. Statistical significance was achieved when the P-value was < 0.05. Kaplan-Meier analysis was used to depict OS.
3. Results
3.1. Patient characteristics
A total of 135 consecutive AML patients who underwent allogeneic stem cell transplantation after having achieved CR1 (median age: 36 years; 61.5% male) were retrospectively reviewed in this study. 57 of these patients underwent HLA-matched HSCT from sibling donors, 38 underwent HLA-matched HSCT from unrelated donors, and 40 received haploidentical HSCT. The characteristics of these AML patients are summarized in Table 1. All patients achieved neutrophil and platelet engraftment post-transplantation. The OS for all patients was 63.7% (86/135) during 7-years. Patients in the survival group were younger than those in the non-survivor group (P=0.020). No significant differences were found between the survival and non-survival groups regarding patients’ gender, WBC count and BM blasts percentage at diagnosis, French-American-British (FAB) classification, chemotherapy cycles for consolidation pre-transplant, donor sex, female donor to male recipient, type of transplantation, graft source, CD34+ cell count, absolute nucleated cell (ANC) count, and CMV/EBV viremia.
3.2 Gene mutations
The profile of gene mutations in these AML patients is shown in Table 2. Gene mutations with incidence > 10% include NRAS, ASXL1, TET2, IDH1/2, C-KIT, CEBPA biallelic, WT1, NPM1, DNMT3A, and FLT3-ITD. Mutational rate of KRAS, RUNX1, GATA2, PTPN11 and TP53 is 7.4%, 6.7%, 7.4%, 7.4% and 0.7%, respectively. The non-survivor group exhibited a higher presence of IDH 1/2 mutation than in the survivor group (20.4% vs. 8.1%, respectively; P=0.039). The non-survivor group also existed trends of a higher rate of mutations in ASXL1, TET2, and FLT3-ITD than the survivor group; although they were not statistically significant, which is likely due to insufficient patients in this study.
|
Characteristics |
Non-survivors (n=49, 36.3%) |
Survivors (n=86, 63.7%) |
Total (n=135) |
P value |
|
Age (years), median (range) |
40(12-58) |
34(10-58) |
36 (10-58) |
0.020 |
|
Male (%) |
33(67.3) |
50(58.1) |
83 (61.5) |
0.290 |
|
WBC count at diagnosis, ×109/L |
32.1(1.6-307.7) |
30.9(1.1-346.9) |
31.3(1.1-346.9) |
0.418 |
|
Bone marrow blasts percentage at diagnosis (%) |
54(5-98) |
69(11.5-97.8) |
62.5(5-98) |
0.053 |
|
FAB type |
- |
- |
- |
0.094 |
|
M0 |
3(6.1) |
1(1.2) |
4(3.0) |
- |
|
M1 |
8(16.3) |
20(23.3) |
28(20.7) |
- |
|
M2 |
16(32.7) |
32(37.2) |
48(35.6) |
- |
|
M4 |
5(10.2) |
15(17.4) |
20(14.8) |
- |
|
M5 |
12(24.5) |
17(19.8) |
29(21.5) |
- |
|
M6 |
4(8.2) |
1(1.2) |
5(3.7) |
- |
|
M7 |
1(2.0) |
0 |
1(0.7) |
- |
|
Risk group at diagnosis |
- |
- |
- |
0.010 |
|
Favorable |
11(22.4) |
29(33.7) |
40(29.6) |
- |
|
Intermediate |
17(34.7) |
41(47.7) |
58(42.9) |
- |
|
Poor |
21(42.9) |
16(18.6) |
37(27.4) |
- |
|
No. of courses to achieve CR1 |
- |
- |
- |
0.043 |
|
1 |
33(67.3) |
71(82.6) |
104(77.0) |
- |
|
>1 |
17(34.6) |
14(16.3) |
31(23.0) |
- |
|
Cycles for consolidation |
- |
- |
- |
0.833 |
|
1 |
27(55.1) |
50(58.1) |
77(57.0) |
- |
|
>1 |
12(24.5) |
22(25.6) |
34(25.2) |
- |
|
Donor sex |
- |
- |
- |
0.960 |
|
Male |
27(55.1) |
47(54.7) |
74(54.8) |
- |
|
Female |
22(44.9) |
39(45.3) |
61(45.2) |
- |
|
Female donor to male recipient |
14(28.6) |
21(24.4) |
35(25.9) |
0.596 |
|
Type of transplantation |
- |
- |
- |
0.968 |
|
HLA-matched sibling donor |
20(40.8) |
37(43.0) |
57(42.2) |
- |
|
HLA-matched unrelated donor |
14(28.6) |
24(27.9) |
38(28.1) |
- |
|
HLA haploidentical donor |
15(30.6) |
25(29.1) |
40(29.6) |
- |
|
Graft |
- |
- |
0.831 |
|
|
BM |
7(14.3) |
14(16.3) |
21(15.6) |
- |
|
PBSCs |
29(59.2) |
52(60.5) |
81(60.0) |
- |
|
BM and PBSCs |
13(26.5) |
20(23.3) |
33(24.4) |
- |
|
Median CD34+ cell counts, ×106/kg |
3.6(1.3-8.1) |
3.6(1.2-9.9) |
3.6(1.2-9.9) |
0.664 |
|
Median ANC counts, ×108/kg |
8.7(3.0-28.5) |
8.3(1.3-18.9) |
8.6(1.3-28.5) |
0.260 |
|
aGVHD grade III- IV |
16(32.7) |
8(9.3) |
24(17.8) |
<0.001 |
|
cGVHD |
17(34.7) |
55(64.0) |
72(53.3) |
<0.001 |
|
Relapse |
27(55.1) |
6(7.0) |
33(24.4) |
<0.001 |
|
CMV |
22(44.9) |
28(32.6) |
50(37.0) |
0.153 |
|
EBV |
9(18.4) |
21(24.4) |
30(22.2) |
0.416 |
Table 1: Characteristics of AML patients undergoing allogeneic HSCT in CR1.
|
Gene mutations |
Non-survivors (n=49, 36.3%) |
Survivors (n=86, 63.7%) |
Total (n=135) |
P value |
|
Signal genes |
||||
|
KRAS |
4(8.2) |
6(7.0) |
10(7.4) |
0.800 |
|
NRAS |
7(14.3) |
17(19.8) |
24(17.8) |
0.423 |
|
C-KIT |
6(12.2) |
18(20.9) |
24(17.8) |
0.204 |
|
FLT3-ITD |
8(16.3) |
9(10.5) |
17(12.6) |
0.324 |
|
PTPN11 |
4(8.2) |
6(7.0) |
10(7.4) |
0.800 |
|
Myeloid transcription factor mutations |
||||
|
CEBPA biallelic |
6(12.2) |
16(18.6) |
22(16.3) |
0.215 |
|
RUNX1 |
4(8.2) |
5(5.8) |
9(6.7) |
0.599 |
|
GATA2 |
3(6.1) |
7(8.1) |
10(7.4) |
0.667 |
|
Chromatin-modifying genes |
||||
|
ASXL1 |
9(18.4) |
10(11.6) |
19(14.1) |
0.279 |
|
EZH2 |
1(2.0) |
5(5.8) |
6(4.4) |
0.306 |
|
DNA methylation-associated genes |
||||
|
TET2 |
9(18.4) |
8(9.3) |
17(12.6) |
0.127 |
|
IDH1/2 |
10(20.4) |
7(8.1) |
17(12.6) |
0.039 |
|
DNMT3A |
6(12.2) |
8(9.3) |
14(10.4) |
0.590 |
|
Tumor suppressor genes |
||||
|
TP53 |
1(2.0) |
0 |
1(0.7) |
0.184 |
|
WT1 |
6(12.2) |
16(18.6) |
22(16.3) |
0.336 |
|
PHF6 |
2(4.1) |
1(1.2) |
3(2.2) |
0.269 |
|
NPM1 |
6(12.2) |
10(11.6) |
16(11.9) |
0.915 |
|
Spliceosome-complex genes |
||||
|
SF3B1 |
1(2.0) |
0 |
1(0.7) |
0.184 |
|
U2AF1 |
1(2.0) |
1(1.2) |
2(1.5) |
0.685 |
|
Cohesin-complex gene |
||||
|
RAD21 |
1(2.0) |
1(1.2) |
2(1.5) |
0.685 |
Table 2: Gene mutations occurred in AML patients.
|
|
Univariate analysis |
Multivariate analysis |
||
|
HR (95% CI) |
P value |
HR (95% CI) |
P value |
|
|
Age |
1.030 (1.004-1.056) |
0.022 |
||
|
Risk group at diagnosis |
||||
|
Favorable |
0.004 |
|||
|
Intermediate |
1.143(0.535-2.441) |
0.730 |
||
|
Poor |
2.815(1.355-5.846) |
0.006 |
||
|
>1 courses to achieve CR1 |
1.923(1.058-3.497) |
0.032 |
||
|
aGVHD III- IV |
3.356(1.838-6.098) |
<0.001 |
2.315(1.245-4.310) |
0.008 |
|
cGVHD |
0.327(0.181-0.591) |
<0.001 |
0.375(0.683-0.206) |
0.001 |
|
Relapse |
4.184(2.370-7.407) |
<0.001 |
3.876(2.174-6.944) |
<0.001 |
|
IDH1/2 |
1.980(0.987-3.984) |
0.054 |
||
HR hazard ratio; CI confidence interval
Table 3: Univariate and multivariate analysis of risk factors for OS in AML patients undergoing allogeneic SCT in CR1.
3.3 Risk factors associated with OS
As shown in Table 1, the percentage of cytogenetic high-risk patients at diagnosis was more frequent in the non-survivor group than the survivor group (42.9% vs. 18.6%); percentage of favorite-risk patients was less common in the non-survivor group (22.4% vs. 33.7%; P=0.010). Seventeen patients (34.6%) required more than one cycle of induction chemotherapy to achieve CR in the non-survivor group, whereas only 14 (16.3%) patients needed twice or more cycles of induction chemotherapies (P=0.043) in survival patients. More patients had aGVHD grade III- IV in the non-survivor group (n=16, 32.7%), comparing with the survivor group (n=8, 9.3%) (P < 0.001). Chronic GVHD was more frequent in the survivor group (64.0% vs. 34.7%; P < 0.001). In addition, patients in the non-survivor group experienced leukemia relapse more frequently comparing with the survivor group (55.1% vs. 7.0%; P < 0.001). Multivariate analysis shown in Table 3 further identified the independent risk factors associated with mortality in these AML patients undergoing allogeneic HSCT in CR1, including aGVHD grade III-IV (HR=2.315, P=0.008), and relapse (HR=3.876, P < 0.001); meanwhile, cGVHD is a favorable predictor (HR=0.375, P=0.001). The OS was significantly inferior in patients occurring aGVHD grade III-IV comparing with those without aGVHD grade III-IV (33.3% and 70.3%, respectively; P < 0.001; Figure 1a). For patients with cGVHD, the OS was significantly higher than those without cGVHD (76.4% and 49.2%, respectively; P < 0.001; Figure 1b). For patients who experienced leukemia relapse post-transplantation, the OS was 18.2%, which was significantly lower than those without leukemia relapse (78.4%) (P < 0.001; Figure 1c).
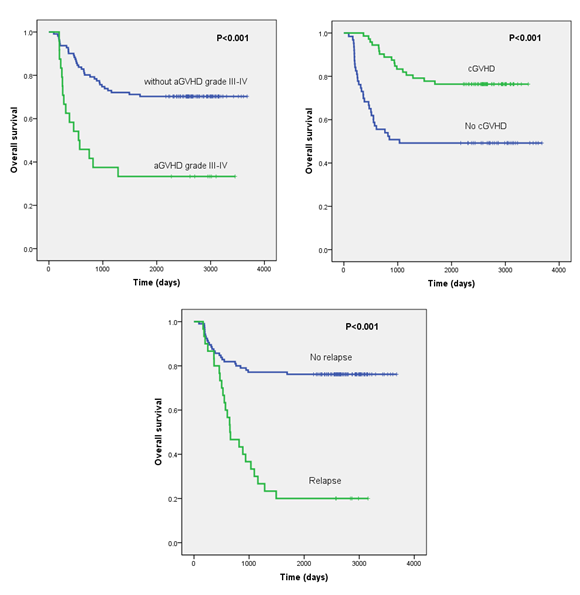
Figure 1: Overall survival (OS) depicted by Kaplan-Meier analysis in AML patients undergoing allogeneic SCT in first complete remission (CR1). (a) OS was significantly lower in patients with acute graft versus host disease (aGVHD) III- IV. (b) OS was significantly lower in patients without chronic graft versus host disease (cGVHD). (c) OS was significantly lower in AML patients with leukemia relapse post-transplantation.
4. Discussion
In this study, we summarized the outcome of AML patients undergoing allogeneic HSCT in CR1, compared the clinical characteristics and gene mutation status at diagnosis between the non-survivor and survivor groups. The OS of all enrolled patients is 63.7% during 7-years, suggesting that allogeneic HSCT represents a potentially curative therapeutic approach for AML patients in CR1. In addition, we identified independent risk factors of OS in allogeneic transplanted AML patients. Acute GVHD grade III-IV and relapse are high adverse risk factors, whereas cGVHD is a protective factor for transplanted AML patients.
Acute GVHD grade III- IV had a significant negative impact on OS in our study. Among the patients (n=24) who experience aGVHD grade III- IV, 66.7% (n=16) of patients died; whereas among those (n=111) who didn’t have aGVHD grade III- IV, 70.3% of patients survived (Table 1). This highlights that aGVHD grade III- IV is a common cause of morbidity and mortality after allogeneic HSCT. The importance of effective control of severe aGVHD should be addressed. MAGIC algorithm probability (MAP) has been validated to estimate the risk of NRM due to aGVHD by adopting two GI biomarkers (ST2 and REG3α) as a “liquid biopsy” of gastrointestinal (GI) crypt damage [7]. Besides, the expression profiles of microRNAs are also investigated to identify organ-specific diagnosis [8]. These strategies help to early detect the presence of aGVHD. A a calcineurin inhibitor (e.g., cyclosporine or tacrolimus) plus an antimetabolite is recommended for GVHD prophylaxis in patients receiving allogeneic HSCT. Cyclosporine serum concentration should be carefully monitored after transplantation and be tapered to balance the risk of aGVHD and relapse. In addition, rabbit ATG (rATG) is also recommended for patients receiving matched unrelated donor HSCT [9].
First-line treatment for aGVHD is corticosteroid. However, only about 50% of patients are effective on corticosteroid therapy [10]. Refractory aGVHD is a challenge to handle. There is no consensus on second-line treatment options. Recently, ruxolitinib was reported to induce an overall response rate of 69.5% and a CR rate of 21.7% in heavily pretreated refractory aGVHD [11]. Adoptive infusion of Treg cells has also been applied to restrain immune responses towards hosts to prevent and treat aGVHD in both pre-clinical and clinical settings [12]. Other available options include basiliximab, infliximab, etanercept, daclizumab, sirolimus, pentostatin, alemtuzumab, rATG, anti-CD3/CD7 immunotoxin, MMF, MTX, vedolizumab, mesenchymal stem cells (MSCs), and fecal microbiota transplantation (FMT) [13, 14].
Our data found that cGVHD is a favorable predictor of OS in transplanted AML patients. The OS of patients with cGVHD and without cGVHD was 76.4% and 49.2%, respectively (P < 0.001). This phenomenon may be explained by the less relapse rate between the two groups (20.8% vs. 28.6%), which implies the more presence of graft versus leukemia (GVL) effect in patients with cGVHD. However, two patients with pulmonary cGVHD and respiratory insufficiency died of respiratory failure. Severe cGVHD increases mortality and reduces patients’ quality of life (QOL), thus special management should be given to these patients to avoid life-threatening consequences.
Leukemia relapse is another significantly unfavorable predictor of AML patients after transplant in CR1. 30% to 80% of AML patients relapse within 2 years after achieving CR1, representing the leading cause of mortality in AML [15]. Up to 50% of AML patients relapse after allogeneic HSCT depending on disease characteristics [16]. They generally follow a dismal prognosis, usually with an OS less than 20% [17]. The relapse rate we reported here is 24.4%. The OS for relapsed patients in our study is 18.2%, which is extremely lower than the non-relapsed AML patients (78.4%) (P < 0.001; Picture 1c). The median interval between allogeneic HSCT and relapse is 331 (30-1101) days. The median interval between relapse and death in the non-survivor group is 123 (19-822) days. This indicates that methods of prevention, early detection, and novel therapies on relapse are particularly important to improve these patients’ outcomes.
MRD is an increasingly important indicator in AML for the early detection of relapse. It should be carefully monitored as deep as possible during and after therapy (e.g., after each cycle of chemotherapy, pre-transplant, and post-transplant period). A positive MRD measured by flow cytometry (FCM) before transplant and non-remission status of gene mutation increases the risk of relapse after HSCT [18]. Furthermore, it can be the early molecular sign of relapse to trigger pre-emptive interventions prior to an imminent hematological relapse. Approaches for MRD detection can be achieved by monitoring mutated genes (e.g., FLT3-ITD), distinct fusion gene transcripts (e.g., RUNX1-RUNX1T1, CBFB-MYH11) and overexpressed genes like WT1 using sequencing technique or real-time quantitative polymerase chain reaction (RQ-PCR) [19-22]. Chimerism analysis using STR-PCR or digital PCR (dPCR), multiparameter flow cytometry (MFC) and fluorescence in situ hybridization (FISH) are also practical methods for MRD detection [23,24]. In the setting of overt relapse, formidable challenges exist. Post-transplanted patients usually can’t tolerate intensive chemotherapy. They are often refractory to low-dose chemotherapy. And remission post-chemotherapy is usually transient. Donor lymphocyte infusion (DLI) is effective in a small fraction of relapse cases [17]. A second allogeneic HSCT is unlikely to be tolerated or available in these fragile patients due to prior cytotoxic conditioning regimen. New therapeutic methods are urgently needed to rescue their dismal prognosis.
During the last three years, eight novel drugs were approved by the Food and Drug Administration (FDA) in the AML therapeutic landscape. They are FLT3 inhibitor (midostaurin and gilteritinib), CPX-351 (liposomal cytarabine and daunorubicin), gemtuzumab ozogamicin (anti-CD33 antibody drug conjugate), enasidenib (IDH2 inhibitor), ivosidenib (IDH1 inhibitor), venetoclax (BCL-2 inhibitor), and glasdegib (hedgehog inhibitor) [25, 26]. Other optional agents include azacitidine, decitabine (hypomethylating agents), and panobinostat (histone deacetylase inhibitor). These drugs provided both new opportunities and challenges for hematologists. The timing point of applying these drugs as prophylaxis (no present leukemia evidence), pre-emptive approach (at the time of positive MRD), or salvage therapy (overt hematologic relapse), as well as single-use or a combination of other anti-leukemia approaches should be vigorously tested in clinical trials to examine their full potency.
Despite the above-mentioned drugs with direct antileukemic activity, another method is to navigate the immune system. Chimeric antigen receptor (CAR)-modified T cells targeting leukemia antigens, including CD33, CD123, C-type lectin-like molecule-1 (CLL-1), CD7, FLT3, CD13 and TIM3 has been reported [27-31]. However, side effects on normal hematopoiesis suppression, cytokine release syndrome (CRS) and immune escape from the aforementioned CAR-T needs to be delicately resolved. It is well known that the effect of allogeneic HSCT relies on the GVL effect mediated by the rebuilding of the host-derived immune system against malignant cells [32]. Frequent loss of HLA-DR, -DQ and -DP on leukemia cells, and deregulation of multiple costimulatory ligands is documented to be responsible to drive AML relapse post-HSCT. These mechanisms reduce donor T cells’ recognition of leukemic cells. Interferon-γ or immune checkpoint inhibitors are hopeful to offset these detrimental changes, respectively [33]. This report provides new insights into the activation of the immune system to fight against leukemia.
Gene mutation was common in AML patients. In the present study, the IDH1/2 gene was found to be more frequently mutated in the non-survivor group than in the survivor group. Furthermore, the higher relapse rate was found in the mutation group (41.2% vs. 22.0%), which led to higher mortality. This high relapse rate implies allogeneic HSCT itself may not be sufficient to overcome this specific mutation. More cases are needed to confirm our findings. However, due to its relapse tendency, integration of targeted therapy, e.g., IDH2 inhibitor enasidenib and IDH1 inhibitor ivosidenib into the procedure of allogeneic HSCT is of particular interest to be further explored.
5. Conclusions
In conclusion, allogeneic HSCT is a curative way for the majority of AML patients in CR1. Acute GVHD grade III-IV and relapse are independent high-risk factors of OS in AML patients undergoing allogeneic SCT in CR1; whereas cGVHD provides a favorable effect on OS. New measures to predict aGVHD, a more effective way to control severe aGVHD, with the integration of novel anti-leukemia agents and exploitation of our immune system to eradicate leukemic cells, is of immense interest to optimize the transplantation outcomes.
Author Contributions
Haiyan Bao- conceptualization, data collection, methodology, writing original draft, review & editing. Vijay Kumar Kolluri- data collection, statistical analysis, editing. Jia Chen- conceptualization, data collection, statistical analysis. Xiao Ma- patients’ management post-transplantation. Yue Han- implement of chemotherapy and stem cell transplantation. Chengcheng Fu- implement of chemotherapy and stem cell transplantation. Mei Yang- conceptualization, methodology, review & editing. Depei Wu- conceptualization, methodology, implement of chemotherapy and stem cell transplant, review & editing.
Funding
This study was supported by the Jiangsu Provincial Key Medical Center (YXZXA2016002), Innovation Capability Development Project of Jiangsu Province (BM2015004), National Key R&D Program of China (2016YFC0902800, 2017YFA0104502, 2017ZX09304021).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Löwenberg B. Acute myeloid leukemia: the challenge of capturing disease variety. Hematology. American Society of Hematology. Education Program (2008): 1-11.
- Singh AK, McGuirk JP. Allogeneic stem cell transplantation: a historical and scientific overview. Cancer research 76 (2016): 6445-6451.
- Bejanyan N, Haddad H, Brunstein C. Alternative donor transplantation for acute myeloid leukemia. Journal of clinical medicine 4 (2015): 1240-1268.
- Bullinger L, Döhner K, Döhner H. Genomics of acute myeloid leukemia diagnosis and pathways. Journal of clinical oncology 35 (2017): 934-946.
- Martino R, Romero P, Subirá M, et al. Comparison of the classic Glucksberg criteria and the IBMTR Severity Index for grading acute graft-versus-host disease following HLA-identical sibling stem cell transplantation. International Bone Marrow Transplant Registry. Bone marrow transplant 24 (1999): 283-287.
- Filiovich AH, Weisdorf D, Pavletic S, et al. National Institutes of Health consensus development project on criteria for clinical trials in chronic graft-versus-host disease: I. Diagnosis and Staging Working Group report. Biology of blood and marrow transplantation: journal of the American Society for Blood and Marrow Transplantation 11 (2005): 945-956.
- Srinagesh HK, Levine JE, Ferrara JLM. Biomarkers in acute graft- versus-host disease: new insights. Therapeutic advances in hematology 10 (2019): 2040620719891358.
- Motaei J, Yaghmaie M, Ahmadvand M, et al. MicroRNAs as potential diagnostic, prognostic, and predictive biomarkers for acute graft-versus-host disease. Biology of blood and marrow transplantation: journal of the American Society for Blood and Marrow Transplantation 25 (2019): e375-e386.
- Penack O, Marchetti M, Ruutu T, et al. Prophylaxis and management of graft versus host disease after stem-cell transplantation for haematological malignancies: updated consensus recommendations of the European Society for Blood and Marrow Transplantation. The Lancet. Haematology 7 (2020): e157-e167.
- Bacigalupo A. Management of acute graft-versus-host disease. British journal of haematology 137 (2007): 87-98.
- Escamilla Gómez V, García-Gutiérrez V, López Corral L, et al. Ruxolitinib in refractory acute and chronic graft-versus-host disease: a multicenter survey study. Bone marrow transplantation 55 (2020): 641-648.
- Elias S, Rudensky AY. Therapeutic use of regulatory T cells for graft-versus-host disease. British journal of haematology 187 (2019): 25-38.
- Ali R, Ramdial J, Algaze S, et al. The role of anti-thymocyte globulin or alemtuzumab-based serotherapy in the prophylaxis and management of graft-versus-host disease. Biomedicines 5 (2017): 67.
- Malard F, Huang XJ, Sim JPY. Treatment and unmet needs in steroid-refractory acute graft-versus-host disease. Leukemia 34 (2020): 1229-1240.
- Rollig C, Bornhauser M, Thiede C, et al. Long-term prognosis of acute myeloid leukemia according to the new genetic risk classification of the European LeukemiaNet recommendations: evaluation of the proposed reporting system. Journal of clinical oncology 29 (2011): 2758-2765.
- De Lima M, Porter DL, Battiwalla M, et al. Proceedings from the National Cancer Institute’s Second International Workshop on the biology, prevention, and treatment of relapse after hematopoietic stem cell transplantation: Part III. Prevention and treatment of relapse after allogeneic transplantation. Biology of blood and marrow transplantation: journal of the American Society for Blood and Marrow Transplantation 20 (2014): 4-13.
- Schmid C, Labopin M, Nagler A, et al. Donor lymphocyte infusion in the treatment of first hematological relapse after allogeneic stem-cell transplantation in adults with acute myeloid leukemia: a retrospective risk factors analysis and comparison with other strategies by the EBMT Acute Leukemia Working Party. Journal of clinical oncology 25 (2007): 4938-4945.
- Chen J, Yang L, Fan Y, et al. Comparison of autologous stem cell transplantation versus haploidentical donor stem cell transplantation for favorable and intermediate-risk acute myeloid leukemia patients in first complete remission. Biology of blood and marrow transplantation: journal of the American Society for Blood and Marrow Transplantation 24 (2018): 779-788.
- Cloos J, Goemans BF, Hess CJ, et al. Stability and prognostic influence of FLT3 mutations in paired initial and relapsed AML samples. Leukemia 20 (2006): 1217-1220.
- Jongen-Lavrencic M, Grob T, Hanekamp D, et al. Molecular minimal residual disease in acute myeloid leukemia. The New England journal of medicine 378 (2018): 1189-1199.
- Tang FF, Xu LP, Zhang XH, et al. Monitoring of post-transplant CBFB-MYH11 as minimal residual disease, rather than KIT mutations, can predict relapse after allogeneic haematopoietic cell transplantation in adults with inv(16) acute myeloid leukaemia. British journal of haematology 180 (2018): 448-451.
- Pozzi S, Geroldi S, Tedone E, et al. Leukaemia relapse after allogeneic transplants for acute myeloid leukaemia: predictive role of WT1 expression. British journal of haematology 160 (2013): 503-509.
- Stahl T, Rothe C, Böhme MU, et al. Digital PCR panel for sensitive hematopoietic chimerism quantification after allogeneic stem cell transplantation. International journal of molecular science 17 (2016): 1515.
- Schuurhuis GJ, Heuser M, Freeman S, et al. Minimal/measurable residual disease in AML: a consensus document from the European LeukemiaNet MRD Working Party. Blood 131 (2018): 1275-1291.
- Rautenberg C, Germing U, Haas R, et al. Relapse of acute myeloid leukemia after allogeneic stem cell transplantation: prevention, detection, and treatment. International journal of molecular science 20 (2019): 228.
- Tiong IS, Wei AH. New drugs creating new challenges in acute myeloid leukemia. Genes, chromosomes & cancer 58 (2019): 903-914.
- Wang J, Chen S, Xiao W, et al. CAR-T cells targeting CLL-1 as an approach to treat acute myeloid leukemia. Journal of hematology and oncology 11 (2018): 7.
- Gomes-Silva D, Atilla E, Atilla PA, et al. CD7 CAR T cells for the therapy of acute myeloid leukemia. Molecular therapy: the journal of the American Society of Gene Therapy 27 (2019): 272-280.
- He X, Feng Z, Ma J, et al. Bispecific and split CAR T cells targeting CD13 and TIM3 eradicate acute myeloid leukemia. Blood 135 (2020): 713-723.
- Jetani H, Garcia-Cadenas I, Nerreter T, et al. CAR T-cells targeting FLT3 have potent activity against FLT3(-)ITD(+) AML and act synergistically with the FLT3-inhibitor crenolanib. Leukemia 32 (2018): 1168-1179.
- Cummins KD, Gill S. Anti-CD123 chimeric antigen receptor T-cells (CART): an evolving treatment strategy for hematological malignancies, and a potential ace-in-the-hole against antigen-negative relapse. Leukemia & lymphoma 59 (2018): 1539-1553.
- Kolb HJ. Graft-versus-leukemia effects of transplantation and donor lymphocytes. Blood 112 (2008): 4371-4383.
- Toffalori C, Zito L, Gambacorta V, et al. Immune signature drives leukemia escape and relapse after hematopoietic cell transplantation. Nature medicine 25 (2019): 603-611.


 Impact Factor: * 4.1
Impact Factor: * 4.1 CiteScore: 2.9
CiteScore: 2.9  Acceptance Rate: 11.01%
Acceptance Rate: 11.01%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 