The Landscape and Prognostic Value of m6A Methylation-related Genes in Low Grade Glioma
Article Information
Lixin Ma1#, Zhihui Liu2#, Hongwei Zhang1, Weihai Ning1, Yanming Qu1, Chunjiang Yu1*
1Department of Neurosurgery, Sanbo Brain Hospital, Capital Medical University, Beijing, 100093, P. R. China
2Department of Gynaecology and Obstetrics, Binzhou Medical University Hospital, Binzhou City, Shandong Province, 256603, P. R. China
*Corresponding Author: Chunjiang Yu, Department of Neurosurgery, Sanbo Brain Hospital, Capital Medical University, Beijing, 100093, P. R. China
Received: 06 April 2021; Accepted: 17 April 2021; Published: 21 May 2021
Citation:
Lixin Ma, Zhihui Liu, Hongwei Zhang, Weihai Ning, Yanming Qu, Chunjiang Yu. The Landscape and Prognostic Value of m6A Methylation-related Genes in Low Grade Glioma. Archives of Microbiology & Immunology 5 (2021): 214-231.
View / Download Pdf Share at FacebookAbstract
Background: Low-grade glioma (LGG) can behave aggressively, akin to glioblastoma, and prognostic classification is urgently needed. N6-methyladenosine (m6A) modification is a key regulator of transcriptional expression during tumorigenesis and progression. This study aimed to identify transcriptome biomarkers with prognostic predictive value and define molecular subclassifications.
Methods: We selected 21 m6A methylation-related genes for analysis of 529 LGG samples from TCGA LGG datasets and 1,152 brain tissues from the GTEx datasets. Through difference analysis, Protein-protein interactions (PPI) network, and spearman correlation analysis, gene expression and correlation were studied. Consensus cluster, gene ontology (GO) analysis, Kyoto Encyclopedia of Genes, and Genomes (KEGG) analysis were performed for classification and functional analysis. Lasso Cox regression algorithm and univariate and multivariate analyses were used for assessing risk factors.
Results: The expression of m6A methylation-related genes between normal brain and LGG samples was significantly different. Consensus cluster analysis clearly divided LGG samples into two categories, with a p-value for the difference between prognosis close to 0. Through the lasso Cox regression algorithm and univariate and multivariate analyses, four genetic biomarkers (IGF2BP2, IGF2BP3, YTHDC1, and ALKBH3) were screened out, and the cumulative analysis of these effectively predicted patients’ prognosis.
Conclusion: Consensus cluster analysis based on m6A methylation-related genes clearly divided LGG samples into two categories. Moreover, the cumulative analysis of four genetic biomarkers (IGF2BP2, IGF2BP3, YTHDC1, and ALKBH3) effectively predicted prognosis.
Keywords
Low-grade glioma; N6-methyladenosine; Transcriptome biomarkers; Prognosis; TCGA
Low-grade glioma articles; N6-methyladenosine articles; Transcriptome biomarkers articles; Prognosis articles; TCGA articles
Article Details
Introduction
Glioma is one of the most common primary brain tumors in adults, accounting for more than 70% of malignant brain tumors [1]. Through conventional histopathology in conjunction with isocitrate dehydrogenase (IDH) mutation and 1p/19q codeletion, World Health Organization (WHO) grade 2 and 3 gliomas are usually classified as low-grade gliomas [2-4]. However, low-grade glioma (LGG) occurs mainly in young patients between the ages of 20 and 40, eventually developing into a high-grade malignant tumor [5,6]. In recent years, some IDH-wt grade 3 astrocytomas have been identified as worse than IDH-mut glioblastomas (GBM) and are reportedly underdiagnosed [7]. Additionally, some LGGs have been shown to have a poorer prognosis and should therefore be further stratified for prognosis. However, controversies regarding this remain [8]. Therefore, despite the same integrated diagnosis via histological type and molecular classification, the heterogeneity in clinical behavior and prognosis has been distinguished and become one of the main challenges in clinical practice [9].
RNA methylation, especially N6-methyladenosine (m6A), is the most important epigenetic regulation after DNA methylation and a key regulator of transcriptional expression [10]. This new regulatory layer is called "epitranscriptomics"[11]. N6-methyladenosine not only occurs in mRNA but also occurs in various tRNAs, rRNAs, long-stranded non-coding RNAs, about 25% of the genome-level transcripts [12]. Through enrichment around stop codons, within 5’-, 3’-untranslated regions, and long internal exons, N6-methyladenosine can regulate RNA splicing [13], translocation, stability [14-16], protein translation [17-19], proliferation, differentiation, and survival. During tumor initiation and progression, N6-methyladenosine can regulate corresponding gene expression by modifying oncogenes and tumor suppressor genes to achieve tumor regulation [20].
Like DNA methylation, m6A modification does not alter the original base sequence and is catalyzed by methyltransferase, demethylase, and m6A-binding proteins. m6A methyltransferase is a complex of multiple genes, such as METTL3, METTL14, WTAP, METTL16, KIAA1429 (VIRMA), RBM15,[21] RBM15B,[22] and ZC3H13, which are called “writers”[23,24]. As RNA methylation is dynamic and reversible, m6A demethylases (“erasers”), such as FTO, ALKBH5, and ALKBH3, can revert the methylated groups in RNA and its coding genes [25]. In addition, m6A modification achieves a variety of biological functions through m6A-binding proteins, identified as variable “readers”. These proteins are diverse, including YTH domain family proteins (YTHDC1, YTHDC2, YTHDF1, YTHDF2, and YTHDF3), [26] insulin-like growth factor 2 (IGF2BPs containing IGF2BP1, IGF2BP2, and IGF2BP3), [27] and the HNRNP family (HNRNP C and HNRNP A2/B1) [28]. Thus, in-depth study of the m6A RNA methylation-related genes above has greatly improved our understanding of the function and mechanisms of m6A modification in LGG occurrence and development [29-32].
In this bioinformatics analysis, we comprehensively analyzed the gene expression RNA-seq data of the 21 m6A RNA methylation-related genes above from 11 GTEx normal brain datasets (n = 1152) and 14 GDC TCGA Low Grade Glioma (LGG) datasets (n = 529). We then attempted to identify new strategies for stratifying the prognosis of LGG.
Materials and Methods
Data collection and preprocessing
The gene expression RNA-seq data and corresponding clinical phenotype were downloaded from 14 GDC TCGA Lower Grade Glioma (LGG) datasets and 11 GTEx normal tissue datasets. All of the transcript RNA-seq datasets were obtained from UCSC Xena (https://xena.ucsc.edu/). UCSC TOIL recomputed and combined the datasets to correct for batch effects and to allow for sample merging. In the TCGA-LGG datasets, data from 529 samples was combined into a genomic Matrix and then log2 (x + 1) transformed. In the GTEx datasets, all of the data were log2 (x + 0.001) transformed and combined. Then, GTEx datasets were retransformed by log2 (x + 1), and 1,152 normal brain tissues were extracted from the GTEx datasets. Finally, TCGA LGG datasets and GTEx normal brain datasets were normalized and merged into one Matrix that included all of the gene expression RNA-seq data from the 1,681 samples.
The clinical data Downloaded from GDC TCGA-LGG and GTEX phenotype. Among 529 LGG patients, prior to resection, only 1 case accepted pharmaceutical treatment and 1 case accepted radiation, and the other 527 specimens were collected before neoadjuvant treatment. Because of GTEX phenotype including sample number, primary site, body site detail, and gender, we only statistically analyzed gender in normal brain tissues and LGG.
Selection of m6A RNA Methylation Regulators
Based on the literature search, 21 related genes were selected from the m6A RNA methylation regulators. The gene expression RNA-seq data, including normal brain and LGG samples, was extracted from the above matrix and reserved for subsequent bioinformatics analysis.
Bioinformatic Analysis
The differential analysis of 1,152 normal brain tissues and 529 LGG samples was run using the ‘Limma’ package from open source software for bioinformatics-Bioconductor for R v3.6.2. In the results, a p-value < 0.001, p-value < 0.01, and p-value < 0.05 are indicated by the symbols "***," "**," and "*," respectively. The significantly differentially expressed genes (DEGs) were visualized through heatmap and vioplot. The Protein-Protein Interaction (PPI) network of m6A RNA methylation related-genes was analyzed using the STRING database (https://string-db.org/), which collect and collate databases on protein-protein interactions and can predict protein-protein relationships, with a required composite score greater than 0.7. The correlation of the m6A RNA methylation related-genes was analyzed using the ‘corrplot’ package from R. Lower grade glioma samples were divided into different groups according to the expression of m6A RNA methylation-related genes through the "Consensus Cluster Plus" package. The different clusters were verified via principal component analysis (PCA) from all genes expressed in the LGG samples. Then, analysis was done using ‘survival’ package and differential analysis, and the results in the different clusters were visualized via heatmap. Through univariate Cox regression analyses and the LASSO (the least absolute shrinkage and selection operator) Cox regression algorithm, 4 genes and their coefficients of the m6A RNA methylation regulators were determined to be related to prognosis. The risk score of every sample was calculated by the sum of every coefficient multiplied by corresponding gene expression. The survival and receiver operating characteristic (ROC) curves were compared by different risk scores. Finally, based on clinical phenotype from the TCGA LGG datasets, univariate and multivariate Cox regression analyses were used to assess the impact of the risk score.
Statistical analysis
The statistical analyses were performed using R v3.6.2 (https://www.r-project.org/) and IBM SPSS 22.0 software. Wilcoxon rank sum test was used to compare the difference in gene expression between normal brain and LGG samples. LGG samples were clustered into two groups and chi-square test was used to analyze the relationship between the clusters and clinical data like age, gender, neoplasm grade, laterality, location, IDH status, and histology. The univariate Cox regression analysis was used to analyze the gene expression of 21 related genes with survival time and status. Chi-square test was also used to analyze the distribution of clinical phenotype to the risk score. The ROC curve was tested for the forecast efficiency of the risk score. The Kaplan–Meier method with a two-sided log-rank test was used to compare the survival time with different groups. The univariate and multivariate Cox regression analyses were used to assess the impact of the risk score and clinical characteristics on prognosis. The risk score for prognosis was calculated according to the following formula:
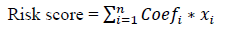
where Coefi is the coefficient, and xi is the relative expression value of z-score transformation of each selected gene. This formula is used to calculate the risk score for each patient in the TCGA LGG data set.
Results
Gene expression RNA-seq of m6A RNA Methylation Regulators in LGG
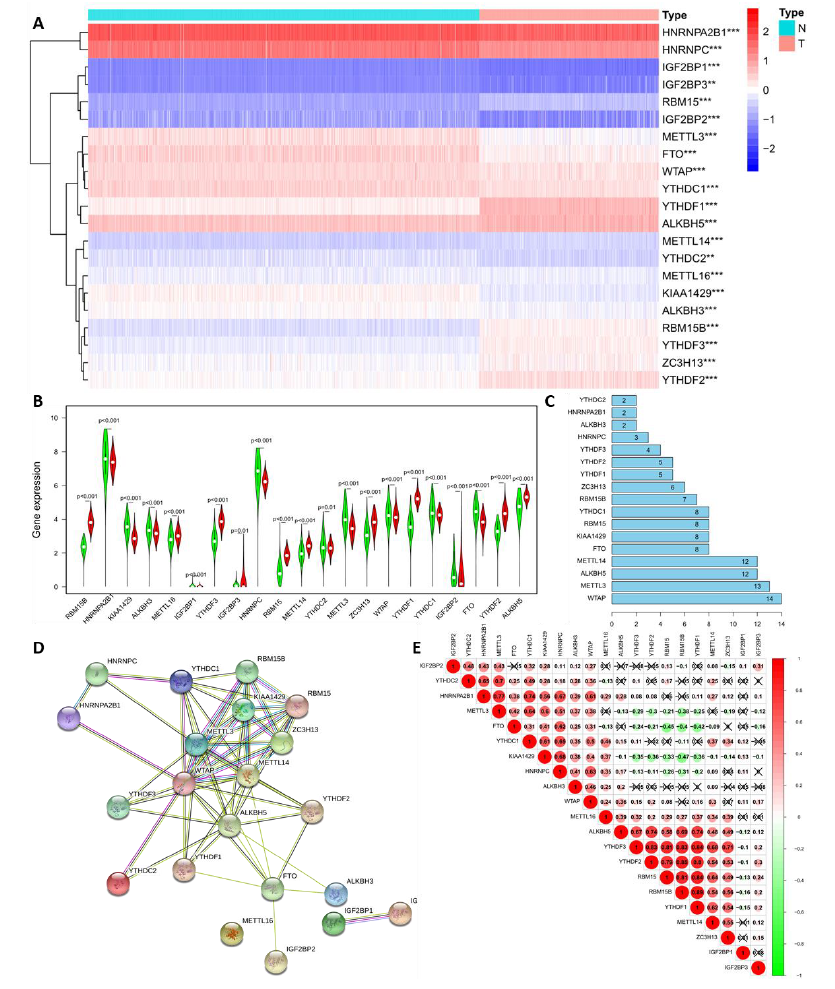
To reveal the important biological functions of m6A RNA Methylation regulators in the formation and progression of LGG, the gene expression RNA-seq data of 21 m6A RNA methylation-related genes were statistically analyzed in 1,152 normal brain tissues from GTEx and 529 LGG cancer samples from TCGA. Compared with normal brain tissues, the difference in the expression of 20 m6A RNA methylation-related genes in LGG samples was highly significant (p < 0.001) where the difference in YTHDC2 gene expression was slightly smaller (P < 0.01; Figure 1A). Among these, the expression of 10 genes (RBM15B, METTL16, YTHDF3, IGF2BP3, RBM15, METTL14, ZC3H13, YTHDF1, YTHDF2, and ALKBH5) increased in the LGG samples, and the expression of others notably decreased (Figure 1B). To better understand the interactions between 21 m6A RNA methylation regulators, PPI network was used to analyze the interactions between these regulators (Figure 1C-D) and spearman’s correlation to analyze the correlation of regulators (Figure 1E). The PPI network can identify proteins with similar functions, known as functional modules, and core genes based on pairwise relationships and nodes. Identifying these core genes is very important for understanding the structure of biological systems. Among the PPI network of 21 m6A RNA methylation-related genes, 61 pairwise relationships were identified. Among them, the hub genes WTAP, METTL3, ALKBH5, METTL14, and FTO had the most connections with other genes (Figure 1C and D), which contained 14, 13, 12, 12 and 8 nodes respectively. To clarify the biological relationship of the above genes, gene correlation was analyzed and is shown in Figure 1E. Like previous research, YTHDF3, YTHDF2, and YTHDF1 in the ‘reader’ and RBM15B and RBM15 in the “writer” groups played synergistic roles. The relationship between ALKBH5 in the “erasers” and YTHDF1 and YTHDF2 in the “readers”; METTL3 in the “writers” and YTHDC2 and HNRNPA2B1 in the readers; RBM15B and RBM15 in the “writers” and YTHDF1 and YTHDF2 in the “readers” were also very related. However, KIAA1429 gene expression was negatively related to YTHDF3, YTHDF2, and YTHDF1 in the ‘readers’ and RBM15B and RBM15 in the “writers.” We speculated that genes at different stages of m6A RNA methylation regulation also had a feedback effect, forming closed-loop biological regulation.
Figure 1: Landscape of m6A RNA methylation-related genes in lower-grade glioma. (A) Gene expression RNA-seq of 21 m6A RNA methylation-related genes in LGG and the difference by statistical analysis. Red is up-regulated and bule is down-regulated in the left diagram. N: normal brain tissue; T: Tumor, which is LGG in the right diagram. (B) Vioplot visualizes gene expression of m6A RNA methylation between normal brain tissue and LGG. Green is normal brain tissue and red is LGG. (C and D) Protein-protein interactions (PPI) network of 21 m6A RNA methylation-related genes in LGG and node analysis of PPI by bar plot. Numbers in Figure C refer to nodes; The circle represents the protein and straight lines represents protein-protein interactions, called pairwise relationships in Figure D. (E) Spearman’s correlation analysis of the 21 m6A modification regulators in LGG. Red is positively correlated and green is negatively correlated. *, **, *** represents p < 0.05, p < 0.01, and p < 0.001, respectively.
LGG consensus clustering based on m6A RNA methylation regulation
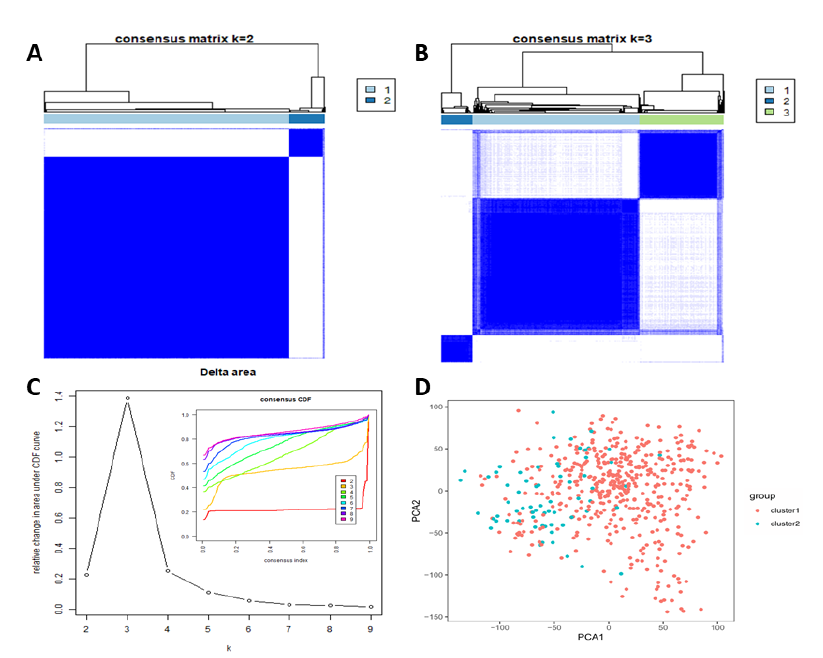
Studies have shown that tumor grade does not fully reflect biological performance, and a new classification was investigated using the ‘Consensus ClusterPlus’ software package. Based on gene expression in the RNA-seq data of 21 m6A RNA methylation-related genes from 529 LGG samples, nine different clusters were tested with different CDF values. Based on consensus cluster analysis research, with the change of K value, the change of CDF value tends to be flat, and its K value can be used as the best cluster number of the data. We found that, when K>4, the relative change of the area under the CDF curve did not change significantly. Therefore, in the TCGA data set, k = 3 seems to have a small CDF value (Figure 2C), but after dividing it into three groups, the correlation between the groups is high (Figure 2B). Therefore, two clusters were identified based on smaller CDF values and correlation between the groups (Figure 2A, 2C). The new classification of two subclasses was further analyzed and verified using PCA of all gene expression (Figure 2D). The results showed that cluster 1 can gathered together and cluster 2 can also be gathered together (Figure 2D), and the results contributed to the next analysis of clinical characteristics.
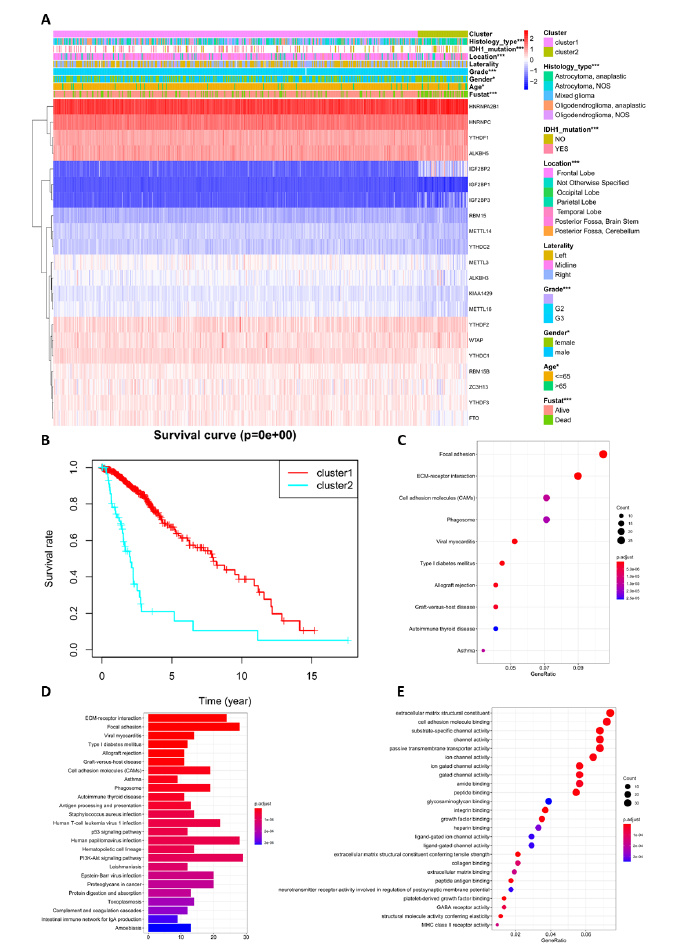
To further verify the impact of new classification on prognosis, the survival curves of the two subclasses were analyzed for 529 LGG patients (Figure 3). The overall survival of cluster 1 and cluster 2 were highly different, with a p-value of 0e + 00, close to zero. Additionally, the 5-year survival rates of clusters 1 and 2 were 66.2% and 15.8%, respectively. This classification predicted prognosis more accurately, superior to tumor grade and histology type which are currently in use. Then, the correlation of the two clusters and their clinical characteristics was explored. Figure 3A shows that the clustering was closely related with information such as tumor grade, IDH mutation, histology type, and location; the correlation p-value was less than 0.001. There was also a correlation with gender and age, but these were not related to tumor laterality. This new classification thoroughly distinguished the malignancy and prognosis of LGG. Then, to elaborate on the clustering results and biological functions of cluster 2, 601 differentially expressed genes (DEGs) were identified by difference analysis between cluster 1 and cluster 2. Gene Ontology (GO) enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis were used to enrich upregulated and downregulated genes in the cluster 2 and cluster 1 subgroups. Differential genes were most enriched in extracellular matrix structural constituent, cell adhesion molecule binding, substrate-specific channel activity, and glycosaminoglycan binding, as shown by GO analysis (Figure 3E). KEGG analysis indicated that these genes play an important role in the PI3K-Akt signaling pathway, focal adhesion, human papillomavirus infection, ECM-receptor interaction, cell adhesion molecules (CAMs), and phagosome. (Figure 3C-D). In short, the classification integrated clinical information and accurately predicted patient prognosis in clinical practice.
Furthermore, Cluster 2 data was further analyzed, including 62 LGG patients. Among them, there were 9 cases of G2 and 53 cases of G3 in tumor grade. In histology type, 38 Astrocytoma Anaplastic, 5 Astrocytoma NOS, 7 Oligodendroglioma Anaplastic, 3 Oligodendroglioma NOS and 8 Mixed gliomas were included. It could be shown that these tumors were mainly anaplastic and NOS, with high degree of malignancy and GBM characteristics.
We also used 21 m6A RNA methylation-related genes to carry out the same consensus cluster for the gene expression RNA-seq data from GDC TCGA GBM datasets mentioned before, and found that the correlation between their clusters was too high to cluster like those of LGG. We concluded that due to the complexity of GBM initiation and development, epigenetic changes such as DNA methylation, mutation, deletion, and amplification may also play an important role in addition to m6A RNA methylation.
Figure 2: Consensus clustering of m6A RNA methylation-related genes. (A) Consensus matrix k = 2; (B) Consensus matrix k = 3; (C) The cumulative distribution function (CDF) of consensus clustering varied with k from two to nine in right diagram; relative change in area under CDF curve with k = 2–9 in left diagram; (D) principal component analysis (PCA) of all gene expression RNA-seq in the TCGA LGG datasets. LGG was divided into two categories: cluster1 (red), cluster2 (blue).
Figure 3: Different clinical characteristics and biological functions of the two clusters. (A) Heatmap shows gene expression RNA-seq and clinical characteristics of the two clusters (cluster 1/2), and statistical analysis of the correlation between the two clusters and clinical characteristics. (B) Kaplan–Meier overall survival (OS) curves of the two clusters from 529 TCGA LGG patients. Cluster 1 subgroup are marked with red, and cluster 2 subgroup are marked with light blue. (C-D) KEGG enrichment analysis for genes with upregulated and downregulated expression between clusters 2 and 1. (E) GO enrichment analysis for genes with upregulation and downregulation of expression between cluster 2 and cluster 1. *, **, *** represents p < 0.05, p < 0.01 and p < 0.001, respectively.
Prognostic Value of the lasso regression risk score
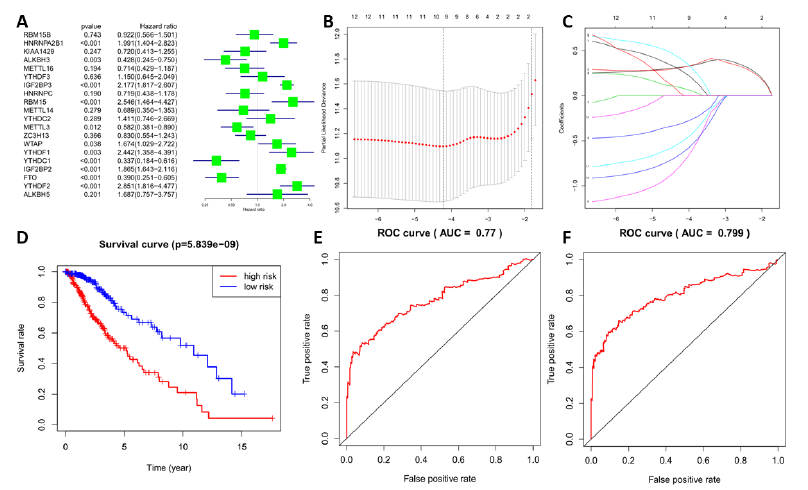
To clarify the effect of m6A RNA methylation-related genes on prognosis in LGG, univariate Cox regression analysis and forest illustration of gene expression were established, and 12 genes related to prognosis were identified (Figure 4A). Eight genes with a high hazard ratio that may shorten the survival time of patients with LGG, such as RBM15 (HR = 2.55, 95% CI = 1.46–4.43), IGF2BP2 (HR = 1.86, 95% CI = 1.64–2.12), IGF2BP3 (HR = 2.18, 95% CI = 1.82–2.61), YTHDF2 (HR = 2.85, 95% CI =1.82–4.48), HNRNPA2B1 (HR = 1.99, 95% CI = 1.40–2.82), YTHDF1 (HR = 2.44, 95% CI = 1.36–4.39), WTAP (HR = 1.67, 95% CI = 1.03–2.72), and IGF2BP1 (HR = 168988.16, 95% CI = 119.95–238073557.7), were screened. Four low risk ratio genes were FTO (HR = 0.39, 95% CI = 0.25–0.60), YTHDC1 (HR = 0.34, 95% CI = 0.1–0.62), ALKBH3 (HR = 0.43, 95% CI = 0.24–0.75), and METTL3 (HR = 0.58, 95% CI = 0.38–0.89). These genes played a positive role in the prognosis of patients with LGG.
To better predict the prognosis of patients with LGG based on the 12 m6A RNA methylation-related genes above, a model was established through lasso regression analysis. According to the lambda value and minimum partial likelihood deviance (Figure 4B and C), 4 genes (IGF2BP2, IGF2BP3, YTHDC1, and ALKBH3) in the LGG datasets were identified, and their coefficients were calculated through lasso regression algorithm. Then, through gene expression and the corresponding coefficient, a risk score for each sample was calculated, and all of the samples in the LGG datasets were divided into a high-risk or a low risk group. The low risk group had better survival than the high-risk group, as shown through Kaplan–Meier overall survival (OS) curves; the differences in OS were statistically significant (p < 0.001; Figure 4D). Then, the risk score was applied to predict a 3-year survival rate via ROC curve. The area under the curve (AUC) was 0.77 (Figure 4E). The results showed that the risk score could estimate the survival time of the patient and accurately predict the patient's prognosis. Furthermore, 10 genes with minimum partial likelihood deviation (IGF2BP2, IGF2BP3, FTO, HNRNPA2B1, YTHDC1, RBM15, IGF2BP1, YTHDF1, ALKBH3, and METTL3) were used to build another model through lasso regression analysis. The AUC of the 10-gene model was 0.799, slightly better than the four-gene model above (Figure 4F).
Figure 4: Establishment of the lasso regression model and the effect of risk score on prognosis (A) The hazard ratios (HR) of m6A RNA methylation-related genes by univariate Cox regression. Hazard ratio (HR), 95% confidence interval (CI) is calculated by univariate Cox regression; The HR of 12 genes was statistically significant and determined for the next analysis; (B, C) Four genes and their corresponding coefficients were screened to construct a risk model by lasso regression analysis. (D) Kaplan–Meier overall survival (OS) curves of high and low risk groups, based on risk score calculated by four genes. (E) ROC curves of prognostic efficiency of risk score calculated by four genes. (F) Prognostic efficiency of risk score calculated by ten genes.
Risk scores accurately predicted patient prognosis
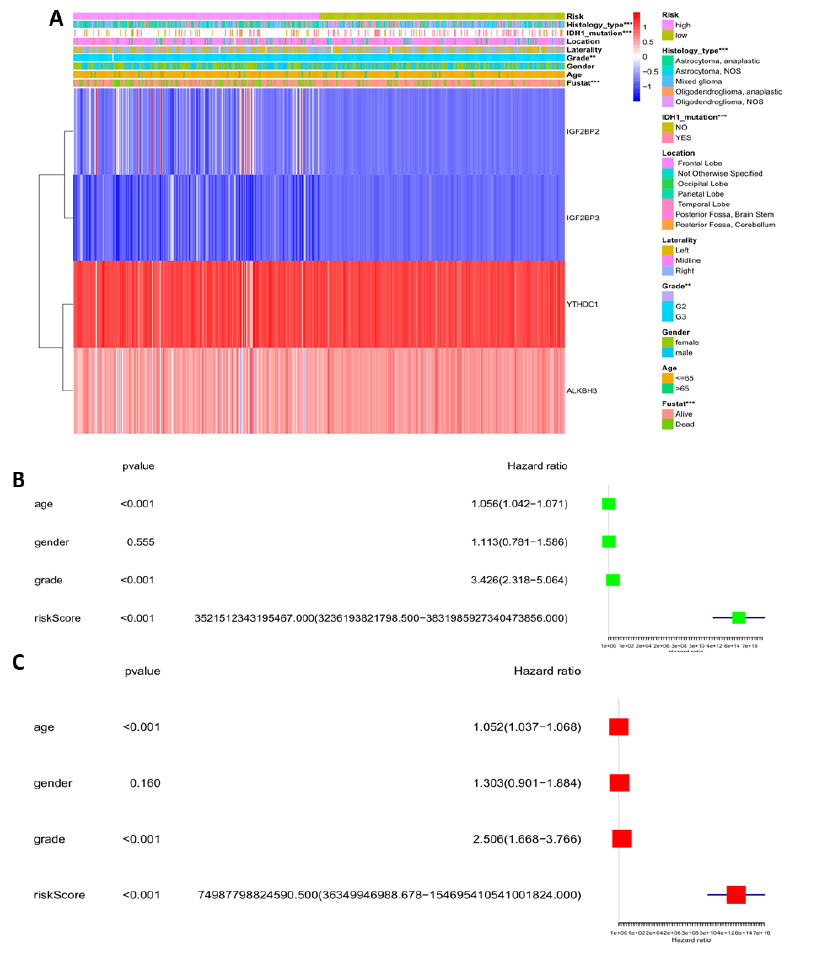
Since risk score was closely related with prognosis, and the relationship between risk score and clinical characteristics may aid in identifying clinical risk factors in clinical practice. Common clinical features from the TCGA LGG datasets such as age, gender, IDH mutation, tumor grade, laterality, location, and histology type were considered. Chi-square test indicated that clinical characteristics such as IDH mutation, tumor grade, and histology type greatly influenced the risk score. Clinical features like age, gender, laterality, and location had no effect on the risk score. Moreover, compared with the low-risk group, the IGF2BP2 and IGF2BP3 genes were up-regulated and the YTHDC1 and ALKBH3 genes were down-regulated in the high-risk group.
To further reveal the impact of clinical features and risk score on prognosis, the univariate and multivariate Cox regression analyses were also discussed. The hazard ratio (HR) provides statistical tests for the efficacy of the risk score and assesses the relative risk for prognosis. Age, tumor grade, and risk score were closely related to the patient's prognosis. This was especially true for the risk score, the hazard ratio (HR) was 3521512343195467.0 in the univariate analysis and 74987798824590.5 in the multivariate analysis. These results indicated that a slight increase in risk score may increase the risk of the patient.
Figure 5: Relationship between the risk score and clinical characteristics, and its impact on prognosis. (A) Heatmap showing gene expression RNA-seq of the four genes screened above and clinical characteristics of the high and low risk LGG patients. Contribution of clinical characteristics to the risk score was statistically analyzed. (B) Univariate Cox regression analyses of the relationship between clinical characteristics, the risk score, and overall survival of patients in the TCGA LGG datasets. (C) Relationship between clinical characteristics, the risk score, and overall survival of patients in the TCGA LGG datasets. *P < 0.05, **P < 0.01, and ***P < 0.001.
Gene expression of four genes screened in human tissues
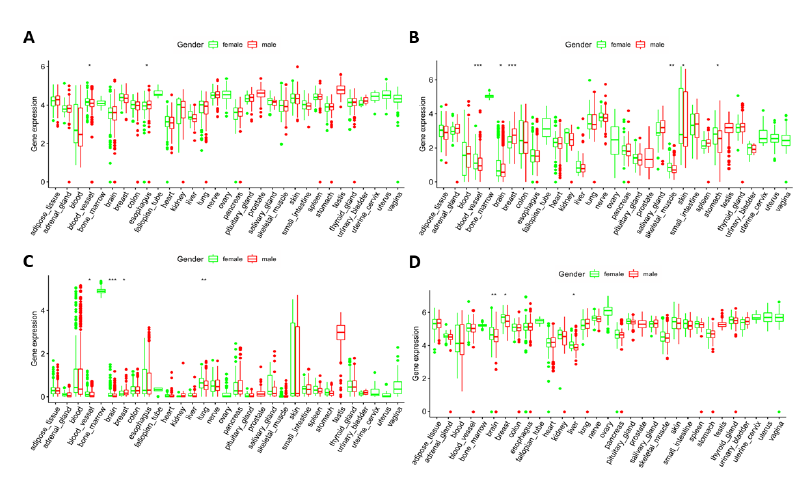
RNA-seq gene expression data of the four genes screened above was studied in human normal tissues. Genotype-Tissue Expression (GTEx) dataset includes genotype data from approximately 714 donors and approximately 11,688 RNA-seq samples from 53 tissue sites and 2 cell lines. The GTEx dataset has sufficient capacity to detect quantitative trait loci in 48 tissues, including 31 solid organ tissues, 10 brain regions, and whole blood. GTEx dataset analysis showed that expression of the IGF2BP2, IGF2BP3, YTHDC1, and ALKBH3 genes was present in 31 human solid organ tissues (Figure 6). In most tissues, expression of the IGF2BP3 gene was low while expression of ALKBH3 and YTHDC1 was high. Based on the hazard ratios (HR) in the univariate Cox regression, we speculated that IGF2BP3 may be the best biomarker.
Discussion
The epigenetic landscape of low grade glioma (LGG), especially crucial genetic alterations, has rapidly changed the understanding of glioma biology and its prognosis [33]. In the revised 2016 WHO Classification of Tumors of the Central Nervous System (CNS), an integrated approach was advocated for clinical practice, including conventional histopathology and genetic features [34]. However, due to the heterogeneity of tumor tissues, some lower-grade gliomas may behave as aggressively as glioblastoma and result in worse outcomes than IDH-mutation GBM [8,9]. Therefore, prognostic classification and development of new strategies for dividing this tumor type into groups with favorable and unfavorable outcomes is needed [35].
With advances in technology, identification of RNA modifications such as transcriptome alterations have attracted the interest of biologists worldwide. m6A modification, a pleiotropic regulator in multiple cancers, occurs specifically at the RRm6AACH motif in mRNA [36]. Like DNA methylation, methyltransferases (writers), demethylases (erasers), and m6A binding proteins (readers) are dynamic during the occurrence and development of cancer [37]. Among these, a balance is needed between methyltransferases and demethylases. The m6A methyltransferase complex consists of METTL3 and METTL14, their adapters WTAP, RBM15, RBM15B, HAKAI, VIRMA (KIAA1429), and ZC3H13, and the methylase METTL16 [38,39]. m6A demethylation is catalyzed by ALKBH5, ALKBH3, and FTO. The balance maintains a dynamic process. RNA alterations after m6A modification require the binding of m6A binding proteins to initiate different downstream effects and exert biological functions [40]. Based on their interaction with m6A in RNA, m6A binding proteins are classified as either direct readers, such as YTH domain containing proteins (YTHDC1-2), the YTH-family proteins (YTHDF1-3), and IGF2BP1-3, or indirect readers, like HNRNP A2/B1 and HNRNP C [41,42]. Therefore, this study analyzed 21 m6A methylation-related genes in 529 LGG samples from the TCGA LGG dataset and 1,152 brain tissue from the GTEx dataset to identify transcriptome markers with prognostic predictive value and define molecular subclassifications.
This study found that compared with normal brain tissues, the expression of m6A methylation-related genes in LGG samples was significantly different, inferring that m6A methylation-related genes may be related to LGG tumorigenesis and progression. In addition, we analyzed the protein-protein interaction (PPI) networks and correlation analysis of 21 m6A methylation-related genes and found that similar to previous studies, some genes like YTHDF3, YTHDF2, YTHDF1 or , RBM15B, and RBM15 played a synergistic role and some genes RBM15, KIAA1429 and YTHDF1 respectively affected writers, erasers, and readers, with a potential feedback effect. Next, using 21 m6A methylation-related genes for consensus cluster analysis, the LGG samples were divided into two categories. The difference in prognosis between the two clusters was highly significant, with a p-value close to 0. The LGG samples were clearly divided into a favorable and an unfavorable group, and this clustering was closely related to clinical features like patient age, gender, IDH mutation, tumor grade, location, and histology type. The difference between clusters 2 and 1 was also analyzed, and 601 differential genes were identified. Through GO enrichment and KEGG pathway analysis, differential genes were enriched in terms of extracellular matrix structural constituent, cell adhesion molecule binding, PI3K-Akt signaling pathway, focal adhesion, and ECM-receptor interaction. These results will facilitate our future research.
We also used the univariate Cox regression analyses to screen out 12 genes related to prognosis and established risk models through the lasso cox regression algorithm. Finally, four genetic biomarkers with prognostic predictive value were selected and the risk score for each sample was calculated. Through survival analysis and ROC analysis, the risk score accurately predicted the patient's prognosis. The relationship between clinical characteristics and risk score was discussed, and the risk score was correlated with IDH mutation, tumor grade, and histology type. Through univariate and multivariate analysis, we found that risk score had the greatest effect on prognosis, indicating that the cumulative analysis of these four genes could effectively predict the prognosis of LGG patients.
In recent years, liquid biopsy, especially in blood, has received considerable attention in monitoring disease response and tracking tumor development. Finding specific molecular markers for early detection and prognosis is a hot spot in glioma research. Through lasso regression algorithm, risk score calculated by 4 genes (IGF2BP2, IGF2BP3, YTHDC1, and ALKBH3) in the LGG datasets could accurately predict the patient's prognosis. Then, the gene expression of these four genetic biomarkers, (IGF2BP2, IGF2BP3, YTHDC1, and ALKBH3) was analyzed in 31 human solid organs. In most tissues, the expression of the IGF2BP3 gene was low while the expression of ALKBH3 and YTHDC1 was high. Based on the hazard ratios (HR) in the univariate Cox regression, we speculated that IGF2BP3 may be the best biomarker in liquid biopsy. Of course, these four genetic biomarkers are merely the results of bioinformatics analysis and need further experimental and clinical validation.
Although RNA methylation in cancer did not attract research until very recently, and no small-molecule inhibitors of RNA methyltransferases are currently available, Small-molecule FTO inhibitors, including rhein, meclofenamic acid 2 (MA2), and R-2-hydroxyglutarat (R-2HG), have been shown to induce growth inhibition and apoptosis in cancer cells, and R-2HG independently or conjunctly with other anticancer drugs can synergisticly block the progression of leukemia in mice [43]. However, we believe that, in the future, effective and specific small-molecule m6A modification inhibitors can be developed through small-molecule compound library screening and/or chemical synthesis, and their pharmacokinetic, safety and anticancer efficacy can be tested in animal models [12].
Ultimately, through the bioinformatic analysis, we found that m6A methylation-related genes play an important role in the occurrence and progression of low-grade glioma. Using consensus cluster analysis, LGG samples were accurately divided into a favorable and an unfavorable group. Next, through the lasso regression algorithm, four genetic biomarkers (GF2BP2, IGF2BP3, YTHDC1, and ALKBH3) were screened out, and the cumulative analysis effectively predicted the prognosis of patients. IGF2BP3 was under-expressed in most normal tissues and had a high hazard ratio, suggesting that it may be the best biomarker in liquid biopsy. However, further research is needed to confirm the clinical significance of genetic biomarkers in LGG. Finally, the understanding of m6A modification is still in its infancy and the application of m6A modification in LGG diagnosis and prognosis needs further research and clinical verification.
Data Availability Statement
Publicly available datasets were analyzed in this study. The datasets can be accessed and downloaded in GDC TCGA (the Cancer Genome Atlas) database (HTSeq-FPKM)]and the Genotype-Tissue Expression (GTEX) [https://xenabrowser.net/datapages (FPKM)].
Acknowledgments
- This work was supported by National Natural Science Foundation of China (No. 22077120) and China Postdoctoral Science Foundation (No. 2019M660715).
- The authors gratefully acknowledge contributions from the TCGA network and the UCSC Network.
- We thank LetPub (www.letpub.com) for its linguistic assistance during the preparation of this manuscript.
Competing interests:
The authors declare no competing interests.
References
- Ostrom QT, Cioffi G, Gittleman H, et al. CBTRUS Statistical Report: Primary Brain and Other Central Nervous System Tumors Diagnosed in the United States in 2012-2016. Neuro Oncology 21 (2019): v1-v1
- Rogers TW, Toor G, Drummond K, et al. The 2016 revision of the WHO Classification of Central Nervous System Tumours: retrospective application to a cohort of diffuse gliomas. Journal of Neuro-Oncology 137 (2018): 181-18
- Cancer Genome Atlas Research Network. Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. New England Journal of Medicine 372 (2015): 2481-24
- Eckel-Passow JE, Lachance DH, Molinaro AM, et al. Glioma groups based on 1p/19q, IDH, and TERT promoter mutations in tumors. New England Journal of Medicine 372 (2015): 2499-2
- Duffau H, Taillandier L. New concepts in the management of diffuse low-grade glioma: proposal of a multistage and individualized therapeutic approach. Neuro-Oncology 17 (2015): 332-3
- Buckner J, Giannini C, Eckel-Passow J, et al. Management of diffuse low-grade gliomas in adults—use of molecular diagnostics. Nature Reviews Neurology 13 (2017):
- Reuss DE, Kratz A, Sahm F, et al. Adult IDH wild type astrocytomas biologically and clinically resolve into other tumor entities. Acta Neuropathologica 130 (2015): 407-4
- Aibaidula A, Chan AK, Shi Z, et al. Adult IDH wild-type lower-grade gliomas should be further stratified. Neuro-Oncology 19 (2017): 1327-13
- Tabouret E, Nguyen AT, Dehais C, et al. Prognostic impact of the 2016 WHO classification of diffuse gliomas in the French POLA cohort. Acta Neuropathologica 132 (2016): 625-6
- Roundtree IA, Evans ME, Pan T, et al. Dynamic RNA modifications in gene expression regulation. Cell 169 (2017): 1187-1
- Hsu PJ, Shi H, He C. Epitranscriptomic influences on development and disease. Genome Biology 18 (2017): 1-9.
- Lan Q, Liu PY, Haase J, et al. The critical role of RNA m6A methylation in cancer. Cancer Research 79 (2019): 1285-12
- Xiao W, Adhikari S, Dahal U, et al. Nuclear m6A reader YTHDC1 regulates mRNA splicing. Molecular Cell 61 (2016): 507-5
- Geula S, Moshitch-Moshkovitz S, Dominissini D, et al. m6A mRNA methylation facilitates resolution of naïve pluripotency toward differentiation. Science 347 (2015): 1002-100
- Knuckles P, Carl SH, Musheev M, et al. RNA fate determination through cotranscriptional adenosine methylation and microprocessor binding. Nature Structural & Molecular Biology 24 (2017):
- Bertero A, Brown S, Madrigal P, et al. The SMAD2/3 interactome reveals that TGFβ controls m 6 A mRNA methylation in pluripotency. Nature 555 (2018): 256-25
- Meyer KD, Patil DP, Zhou J, et al. 5′ UTR m6A promotes cap-independent translation. Cell 163 (2015): 999-1010.
- Wang X, Zhao BS, Roundtree IA, et al. N6-methyladenosine modulates messenger RNA translation efficiency. Cell 161 (2015): 1388-13
- Shi H, Wang X, Lu Z, et al. YTHDF3 facilitates translation and decay of N 6-methyladenosine-modified RNA. Cell Research 27 (2017): 315-3
- Zhao W, Qi X, Liu L, et al. Epigenetic regulation of m6A modifications in human cancer. Molecular Therapy-Nucleic Acids 19 (2020): 405-4
- Pendleton KE, Chen B, Liu K, et al. The U6 snRNA m6A methyltransferase METTL16 regulates SAM synthetase intron retention. Cell 169 (2017): 824-8
- Patil DP, Chen CK, Pickering BF, et al. m 6 A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537 (2016): 369-3
- Wen J, Lv R, Ma H, et al. Zc3h13 regulates nuclear RNA m6A methylation and mouse embryonic stem cell self-renewal. Molecular Cell 69 (2018): 1028-10
- Knuckles P, Lence T, Haussmann IU, et al. Zc3h13/Flacc is required for adenosine methylation by bridging the mRNA-binding factor Rbm15/Spenito to the m6A machinery component Wtap/Fl (2) d. Genes & Development 32 (2018): 415-4
- Ueda Y, Ooshio I, Fusamae Y, et al. AlkB homolog 3-mediated tRNA demethylation promotes protein synthesis in cancer cells. Scientific Reports 7 (2017):
- Meyer KD, Jaffrey SR. Rethinking m6A readers, writers, and erasers. Annual Review of Cell and Developmental Biology 33 (2017): 319-3
- Huang H, Weng H, Sun W, et al. Recognition of RNA N 6-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nature Cell Biology 20 (2018): 285-2
- Alarcón CR, Goodarzi H, Lee H, et al. HNRNPA2B1 is a mediator of m6A-dependent nuclear RNA processing events. Cell 162 (2015): 1299-1
- Liu F, Clark W, Luo G, et al. ALKBH1-Mediated tRNA Demethylation Regulates Translation. Cell 167 (2016):
- Wang S, Sun C, Li J, et al. Roles of RNA methylation by means of N6-methyladenosine (m6A) in human cancers. Cancer Letters 408 (2017): 112-1
- Wang H, Zhu L, Jin H, et al. N6-methyladenosine links RNA metabolism to cancer progression. Cell Death & Disease 9 (2018):
- Wang H, Hu X, Huang M, et al. Mettl3-mediated mRNA m 6 A methylation promotes dendritic cell activation. Nature Communications 10 (2019):
- Gusyatiner O, Hegi ME. Glioma epigenetics: from subclassification to novel treatment options. InSeminars in Cancer Biology 51 (2018): 50-58.
- Louis DN, Perry A, Reifenberger G, et al. The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathologica 131 (2016): 803-8
- Tom MC, Cahill DP, Buckner JC, et al. Management for different glioma subtypes: are all low-grade gliomas created equal?. American Society of Clinical Oncology Educational Book 39 (2019): 133-1
- Liu N, Pan T. N 6-methyladenosine–encoded epitranscriptomics. Nature Structural & Molecular Biology 23 (2016):
- Visvanathan A, Somasundaram K. mRNA Traffic Control Reviewed: N6-Methyladenosine (m6A) Takes the Driver's Seat. Bioessays 40 (2018):
- Warda AS, Kretschmer J, Hackert P, et al. Human METTL16 is a N6-methyladenosine (m6A) methyltransferase that targets pre-mRNAs and various non-coding RNAs. EMBO Reports 18 (2017): 2004-20
- Shima H, Matsumoto M, Ishigami Y, et al. S-Adenosylmethionine synthesis is regulated by selective N6-adenosine methylation and mRNA degradation involving METTL16 and YTHDC1. Cell Reports 21 (2017): 3354-33
- Allis CD, Jenuwein T. The molecular hallmarks of epigenetic control. Nature Reviews Genetics 17 (2016):
- Liu N, Zhou KI, Parisien M, et al. N6-methyladenosine alters RNA structure to regulate binding of a low-complexity protein. Nucleic Acids Research 45 (2017): 6051-60
- Liu N, Dai Q, Zheng G, et al. N 6-methyladenosine-dependent RNA structural switches regulate RNA–protein interactions. Nature 518 (2015): 560-56
- Qiao Y, Zhou B, Zhang M, et al. A novel inhibitor of the obesity-related protein FTO. Biochemistry 55 (2016): 1516-1522.


 Impact Factor: * 3.1
Impact Factor: * 3.1 CiteScore: 2.9
CiteScore: 2.9  Acceptance Rate: 11.01%
Acceptance Rate: 11.01%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 
